+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
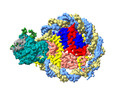
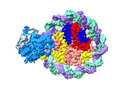
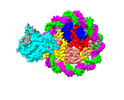
| Title | Structure of DDM1-nucleosome complex in ADP-BeFx state | |||||||||
 Map data Map data | DDM1-nucleosome complex in the ADP-BeFx state | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  complex / complex /  nucleosome / nucleosome /  chromatin remodeling / chromatin remodeling /  structural protein-hydrolase-dna complex / structural protein-hydrolase-dna complex /  GENE REGULATION GENE REGULATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationDNA-mediated transformation /  retrotransposition / chloroplast thylakoid / chromocenter / retrotransposition / chloroplast thylakoid / chromocenter /  thylakoid / response to water deprivation / thylakoid / response to water deprivation /  plasmodesma / plant-type vacuole / DNA methylation-dependent heterochromatin formation / ATP-dependent chromatin remodeler activity ...DNA-mediated transformation / plasmodesma / plant-type vacuole / DNA methylation-dependent heterochromatin formation / ATP-dependent chromatin remodeler activity ...DNA-mediated transformation /  retrotransposition / chloroplast thylakoid / chromocenter / retrotransposition / chloroplast thylakoid / chromocenter /  thylakoid / response to water deprivation / thylakoid / response to water deprivation /  plasmodesma / plant-type vacuole / DNA methylation-dependent heterochromatin formation / ATP-dependent chromatin remodeler activity / plasmodesma / plant-type vacuole / DNA methylation-dependent heterochromatin formation / ATP-dependent chromatin remodeler activity /  plastid / plastid /  chloroplast stroma / heterochromatin formation / epigenetic regulation of gene expression / chloroplast stroma / heterochromatin formation / epigenetic regulation of gene expression /  DNA helicase activity / DNA helicase activity /  chloroplast / response to bacterium / response to wounding / chloroplast / response to bacterium / response to wounding /  peroxisome / structural constituent of chromatin / peroxisome / structural constituent of chromatin /  nucleosome / nucleosome /  DNA helicase / DNA helicase /  chromatin remodeling / protein heterodimerization activity / chromatin remodeling / protein heterodimerization activity /  nucleolus / nucleolus /  ATP hydrolysis activity / ATP hydrolysis activity /  DNA binding / extracellular region / DNA binding / extracellular region /  ATP binding / ATP binding /  nucleus / nucleus /  plasma membrane / plasma membrane /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   Arabidopsis thaliana (thale cress) / synthetic construct (others) Arabidopsis thaliana (thale cress) / synthetic construct (others) | |||||||||
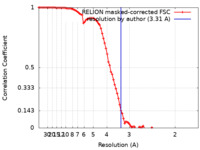
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.31 Å cryo EM / Resolution: 3.31 Å | |||||||||
 Authors Authors | Liu Y / Zhang Z / Du J | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Plants / Year: 2024 Journal: Nat Plants / Year: 2024Title: Molecular basis of chromatin remodelling by DDM1 involved in plant DNA methylation. Authors: Yue Liu / Zhihui Zhang / Hongmiao Hu / Wei Chen / Fan Zhang / Qian Wang / Changshi Wang / Kaige Yan / Jiamu Du /   Abstract: Eukaryotic gene regulation occurs at the chromatin level, which requires changing the chromatin structure by a group of ATP-dependent DNA translocases-namely, the chromatin remodellers. In plants, ...Eukaryotic gene regulation occurs at the chromatin level, which requires changing the chromatin structure by a group of ATP-dependent DNA translocases-namely, the chromatin remodellers. In plants, chromatin remodellers function in various biological processes and possess both conserved and plant-specific components. DECREASE IN DNA METHYLATION 1 (DDM1) is a plant chromatin remodeller that plays a key role in the maintenance DNA methylation. Here we determined the structures of Arabidopsis DDM1 in complex with nucleosome in ADP-BeF-bound, ADP-bound and nucleotide-free conformations. We show that DDM1 specifically recognizes the H4 tail and nucleosomal DNA. The conformational differences between ADP-BeF-bound, ADP-bound and nucleotide-free DDM1 suggest a chromatin remodelling cycle coupled to ATP binding, hydrolysis and ADP release. This, in turn, triggers conformational changes in the DDM1-bound nucleosomal DNA, which alters the nucleosome structure and promotes DNA sliding. Together, our data reveal the molecular basis of chromatin remodelling by DDM1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37535.map.gz emd_37535.map.gz | 6.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37535-v30.xml emd-37535-v30.xml emd-37535.xml emd-37535.xml | 23.5 KB 23.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37535_fsc.xml emd_37535_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_37535.png emd_37535.png | 101 KB | ||
| Filedesc metadata |  emd-37535.cif.gz emd-37535.cif.gz | 7.4 KB | ||
| Others |  emd_37535_half_map_1.map.gz emd_37535_half_map_1.map.gz emd_37535_half_map_2.map.gz emd_37535_half_map_2.map.gz | 49.5 MB 49.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37535 http://ftp.pdbj.org/pub/emdb/structures/EMD-37535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37535 | HTTPS FTP |
-Related structure data
| Related structure data |  8wh9MC  8wh5C  8wh8C  8whaC  8whbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37535.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37535.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
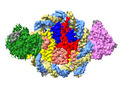
| Annotation | DDM1-nucleosome complex in the ADP-BeFx state | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.855 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half1 map of DDM1-nucleosome complex in the ADP-BeFx state
| File | emd_37535_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half1 map of DDM1-nucleosome complex in the ADP-BeFx state | ||||||||||||
| Projections & Slices |
| ||||||||||||
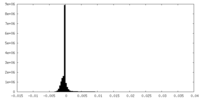
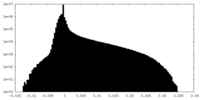
| Density Histograms |
-Half map: Half2 map of DDM1-nucleosome complex in the ADP-BeFx state
| File | emd_37535_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half2 map of DDM1-nucleosome complex in the ADP-BeFx state | ||||||||||||
| Projections & Slices |
| ||||||||||||
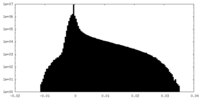
| Density Histograms |
- Sample components
Sample components
+Entire : DDM1-nucleosome complex in ADP-BeFx state
+Supramolecule #1: DDM1-nucleosome complex in ADP-BeFx state
+Macromolecule #1: Histone H3.1
+Macromolecule #2: Histone H4
+Macromolecule #3: Histone H2A.6
+Macromolecule #4: Histone H2B.6
+Macromolecule #7: ATP-dependent DNA helicase DDM1
+Macromolecule #5: DNA (sense strand)
+Macromolecule #6: DNA (antisense strand)
+Macromolecule #8: BERYLLIUM TRIFLUORIDE ION
+Macromolecule #9: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-32 / Number real images: 6776 / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8wh9: |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X