[English] 日本語
 Yorodumi
Yorodumi- EMDB-36333: Atomic structure of wheat ribosome reveals unique features of the... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Atomic structure of wheat ribosome reveals unique features of the plant ribosomes | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Protein Synthesis Machinery /  Eukaryotic Ribosome / Eukaryotic Ribosome /  Plant / Plant /  TRANSLATION / TRANSLATION /  RIBOSOME RIBOSOME | |||||||||
| Biological species |   Triticum aestivum (bread wheat) Triticum aestivum (bread wheat) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.84 Å cryo EM / Resolution: 2.84 Å | |||||||||
 Authors Authors | Mishra RK / Sharma PN / Hussain T | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2024 Journal: Structure / Year: 2024Title: Cryo-EM structure of wheat ribosome reveals unique features of the plant ribosomes. Authors: Rishi Kumar Mishra / Prafful Sharma / Faisal Tarique Khaja / Adwaith B Uday / Tanweer Hussain /  Abstract: Plants being sessile organisms exhibit unique features in ribosomes, which might aid in rapid gene expression and regulation in response to varying environmental conditions. Here, we present high- ...Plants being sessile organisms exhibit unique features in ribosomes, which might aid in rapid gene expression and regulation in response to varying environmental conditions. Here, we present high-resolution structures of the 60S and 80S ribosomes from wheat, a monocot staple crop plant (Triticum aestivum). While plant ribosomes have unique plant-specific rRNA modification (Cm1847) in the peptide exit tunnel (PET), the zinc-finger motif in eL34 is absent, and uL4 is extended, making an exclusive interaction network. We note differences in the eL15-helix 11 (25S) interaction, eL6-ES7 assembly, and certain rRNA chemical modifications between monocot and dicot ribosomes. In eukaryotes, we observe highly conserved rRNA modification (Gm75) in 5.8S rRNA and a flipped base (G1506) in PET. These features are likely involved in sensing or stabilizing nascent chain. Finally, we discuss the importance of the universal conservation of three consecutive rRNA modifications in all ribosomes for their interaction with A-site aminoacyl-tRNA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36333.map.gz emd_36333.map.gz | 141.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36333-v30.xml emd-36333-v30.xml emd-36333.xml emd-36333.xml | 11.8 KB 11.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36333_fsc.xml emd_36333_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_36333.png emd_36333.png | 38.3 KB | ||
| Filedesc metadata |  emd-36333.cif.gz emd-36333.cif.gz | 3.6 KB | ||
| Others |  emd_36333_half_map_1.map.gz emd_36333_half_map_1.map.gz emd_36333_half_map_2.map.gz emd_36333_half_map_2.map.gz | 262.6 MB 262.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36333 http://ftp.pdbj.org/pub/emdb/structures/EMD-36333 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36333 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36333 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36333.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36333.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_36333_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
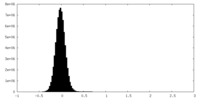
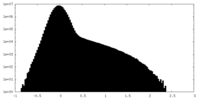
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36333_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
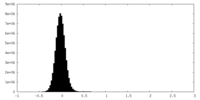
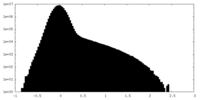
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The small subunit of Ribosome
| Entire | Name: The small subunit of Ribosome |
|---|---|
| Components |
|
-Supramolecule #1: The small subunit of Ribosome
| Supramolecule | Name: The small subunit of Ribosome / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Triticum aestivum (bread wheat) Triticum aestivum (bread wheat) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 75000 / Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 26.54 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z
Z Y
Y X
X