+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tail tip conformation 2 of phage lambda tail | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Bacteriophage / Bacteriophage /  caudovirales / caudovirales /  siphoviridae / siphoviridae /  tail complex / delivery device / macromolecular assembly / tail complex / delivery device / macromolecular assembly /  phage lambda / phage lambda /  cryo-EM / cryo-EM /  VIRAL PROTEIN VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationvirus tail, tube / symbiont genome ejection through host cell envelope, long flexible tail mechanism / viral tail assembly / virus tail / 4 iron, 4 sulfur cluster binding / host cell cytoplasm / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / virion attachment to host cell /  metal ion binding metal ion bindingSimilarity search - Function | |||||||||
| Biological species |   Escherichia phage lambda (virus) Escherichia phage lambda (virus) | |||||||||
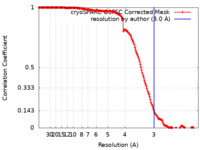
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.0 Å cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Wang JW / Wang C | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2024 Journal: Structure / Year: 2024Title: Architecture of the bacteriophage lambda tail. Authors: Chang Wang / Jinsong Duan / Zhiwei Gu / Xiaofei Ge / Jianwei Zeng / Jiawei Wang /  Abstract: Bacteriophage lambda has a double-stranded DNA genome and a long, flexible, non-contractile tail encoded by a contiguous block of 11 genes downstream of the head genes. The tail allows host ...Bacteriophage lambda has a double-stranded DNA genome and a long, flexible, non-contractile tail encoded by a contiguous block of 11 genes downstream of the head genes. The tail allows host recognition and delivery of viral DNA from the head shell to the cytoplasm of the infected cell. Here, we present a high-resolution structure of the tail complex of bacteriophage lambda determined by cryoelectron microscopy. Most component proteins of the lambda tail were determined at the atomic scale. The structure sheds light on the molecular organization of the extensively studied tail of bacteriophage lambda. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35825.map.gz emd_35825.map.gz | 398.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35825-v30.xml emd-35825-v30.xml emd-35825.xml emd-35825.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35825_fsc.xml emd_35825_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_35825.png emd_35825.png | 66.1 KB | ||
| Filedesc metadata |  emd-35825.cif.gz emd-35825.cif.gz | 7.2 KB | ||
| Others |  emd_35825_half_map_1.map.gz emd_35825_half_map_1.map.gz emd_35825_half_map_2.map.gz emd_35825_half_map_2.map.gz | 392 MB 392 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35825 http://ftp.pdbj.org/pub/emdb/structures/EMD-35825 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35825 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35825 | HTTPS FTP |
-Related structure data
| Related structure data |  8iylMC  8iydC  8iykC  8jvmC  8kgeC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35825.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35825.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.0742 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half 1 map
| File | emd_35825_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half 1 map | ||||||||||||
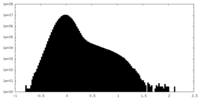
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_35825_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
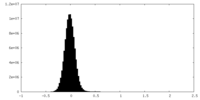
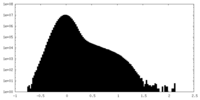
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Tail tip conformation 2 of phage lambda tail
| Entire | Name: Tail tip conformation 2 of phage lambda tail |
|---|---|
| Components |
|
-Supramolecule #1: Tail tip conformation 2 of phage lambda tail
| Supramolecule | Name: Tail tip conformation 2 of phage lambda tail / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:   Escherichia phage lambda (virus) Escherichia phage lambda (virus) |
-Macromolecule #1: Tail tip protein M
| Macromolecule | Name: Tail tip protein M / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia phage lambda (virus) Escherichia phage lambda (virus) |
| Molecular weight | Theoretical: 12.547373 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MKTFRWKVKP GMDVASVPSV RKVRFGDGYS QRAPAGLNAN LKTYSVTLSV PREEATVLES FLEEHGGWKS FLWTPPYEWR QIKVTCAKW SSRVSMLRVE FSAEFEQVVN UniProtKB: Tail tip protein M |
-Macromolecule #2: Tip attachment protein J
| Macromolecule | Name: Tip attachment protein J / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia phage lambda (virus) Escherichia phage lambda (virus) |
| Molecular weight | Theoretical: 124.550625 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MGKGSSKGHT PREAKDNLKS TQLLSVIDAI SEGPIEGPVD GLKSVLLNST PVLDTEGNTN ISGVTVVFRA GEQEQTPPEG FESSGSETV LGTEVKYDTP ITRTITSANI DRLRFTFGVQ ALVETTSKGD RNPSEVRLLV QIQRNGGWVT EKDITIKGKT T SQYLASVV ...String: MGKGSSKGHT PREAKDNLKS TQLLSVIDAI SEGPIEGPVD GLKSVLLNST PVLDTEGNTN ISGVTVVFRA GEQEQTPPEG FESSGSETV LGTEVKYDTP ITRTITSANI DRLRFTFGVQ ALVETTSKGD RNPSEVRLLV QIQRNGGWVT EKDITIKGKT T SQYLASVV MGNLPPRPFN IRMRRMTPDS TTDQLQNKTL WSSYTEIIDV KQCYPNTALV GVQVDSEQFG SQQVSRNYHL RG RILQVPS NYNPQTRQYS GIWDGTFKPA YSNNMAWCLW DMLTHPRYGM GKRLGAADVD KWALYVIGQY CDQSVPDGFG GTE PRITCN AYLTTQRKAW DVLSDFCSAM RCMPVWNGQT LTFVQDRPSD KTWTYNRSNV VMPDDGAPFR YSFSALKDRH NAVE VNWID PNNGWETATE LVEDTQAIAR YGRNVTKMDA FGCTSRGQAH RAGLWLIKTE LLETQTVDFS VGAEGLRHVP GDVIE ICDD DYAGISTGGR VLAVNSQTRT LTLDREITLP SSGTALISLV DGSGNPVSVE VQSVTDGVKV KVSRVPDGVA EYSVWE LKL PTLRQRLFRC VSIRENDDGT YAITAVQHVP EKEAIVDNGA HFDGEQSGTV NGVTPPAVQH LTAEVTADSG EYQVLAR WD TPKVVKGVSF LLRLTVTADD GSERLVSTAR TTETTYRFTQ LALGNYRLTV RAVNAWGQQG DPASVSFRIA APAAPSRI E LTPGYFQITA TPHLAVYDPT VQFEFWFSEK QIADIRQVET STRYLGTALY WIAASINIKP GHDYYFYIRS VNTVGKSAF VEAVGRASDD AEGYLDFFKG KITESHLGKE LLEKVELTED NASRLEEFSK EWKDASDKWN AMWAVKIEQT KDGKHYVAGI GLSMEDTEE GKLSQFLVAA NRIAFIDPAN GNETPMFVAQ GNQIFMNDVF LKRLTAPTIT SGGNPPAFSL TPDGKLTAKN A DISGSVNA NSGTLSNVTI AENCTINGTL RAEKIVGDIV KAASAAFPRQ RESSVDWPSG TRTVTVTDDH PFDRQIVVLP LT FRGSKRT VSGRTTYSMC YLKVLMNGAV IYDGAANEAV QVFSRIVDMP AGRGNVILTF TLTSTRHSAD IPPYTFASDV QVM VIKKQA LGISVV UniProtKB: Tip attachment protein J |
-Macromolecule #3: Tail tip protein L
| Macromolecule | Name: Tail tip protein L / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia phage lambda (virus) Escherichia phage lambda (virus) |
| Molecular weight | Theoretical: 25.730578 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MQDIRQETLN ECTRAEQSAS VVLWEIDLTE VGGERYFFCN EQNEKGEPVT WQGRQYQPYP IQGSGFELNG KGTSTRPTLT VSNLYGMVT GMAEDMQSLV GGTVVRRKVY ARFLDAVNFV NGNSYADPEQ EVISRWRIEQ CSELSAVSAS FVLSTPTETD G AVFPGRIM ...String: MQDIRQETLN ECTRAEQSAS VVLWEIDLTE VGGERYFFCN EQNEKGEPVT WQGRQYQPYP IQGSGFELNG KGTSTRPTLT VSNLYGMVT GMAEDMQSLV GGTVVRRKVY ARFLDAVNFV NGNSYADPEQ EVISRWRIEQ CSELSAVSAS FVLSTPTETD G AVFPGRIM LANTCTWTYR GDECGYSGPA VADEYDQPTS DITKDKCSKC LSGCKFRNNV GNFGGFLSIN KLSQ UniProtKB: Tail tip protein L |
-Macromolecule #4: Tape measure protein
| Macromolecule | Name: Tape measure protein / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia phage lambda (virus) Escherichia phage lambda (virus) |
| Molecular weight | Theoretical: 92.39343 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MAEPVGDLVV DLSLDAARFD EQMARVRRHF SGTESDAKKT AAVVEQSLSR QALAAQKAGI SVGQYKAAMR MLPAQFTDVA TQLAGGQSP WLILLQQGGQ VKDSFGGMIP MFRGLAGAIT LPMVGATSLA VATGALAYAW YQGNSTLSDF NKTLVLSGNQ A GLTADRML ...String: MAEPVGDLVV DLSLDAARFD EQMARVRRHF SGTESDAKKT AAVVEQSLSR QALAAQKAGI SVGQYKAAMR MLPAQFTDVA TQLAGGQSP WLILLQQGGQ VKDSFGGMIP MFRGLAGAIT LPMVGATSLA VATGALAYAW YQGNSTLSDF NKTLVLSGNQ A GLTADRML VLSRAGQAAG LTFNQTSESL SALVKAGVSG EAQIASISQS VARFSSASGV EVDKVAEAFG KLTTDPTSGL TA MARQFHN VSAEQIAYVA QLQRSGDEAG ALQAANEAAT KGFDDQTRRL KENMGTLETW ADRTARAFKS MWDAVLDIGR PDT AQEMLI KAEAAYKKAD DIWNLRKDDY FVNDEARARY WDDREKARLA LEAARKKAEQ QTQQDKNAQQ QSDTEASRLK YTEE AQKAY ERLQTPLEKY TARQEELNKA LKDGKILQAD YNTLMAAAKK DYEATLKKPK QSSVKVSAGD RQEDSAHAAL LTLQA ELRT LEKHAGANEK ISQQRRDLWK AESQFAVLEE AAQRRQLSAQ EKSLLAHKDE TLEYKRQLAA LGDKVTYQER LNALAQ QAD KFAQQQRAKR AAIDAKSRGL TDRQAEREAT EQRLKEQYGD NPLALNNVMS EQKKTWAAED QLRGNWMAGL KSGWSEW EE SATDSMSQVK SAATQTFDGI AQNMAAMLTG SEQNWRSFTR SVLSMMTEIL LKQAMVGIVG SIGSAIGGAV GGGASASG G TAIQAAAAKF HFATGGFTGT GGKYEPAGIV HRGEFVFTKE ATSRIGVGNL YRLMRGYATG GYVGTPGSMA DSRSQASGT FEQNNHVVIN NDGTNGQIGP AALKAVYDMA RKGARDEIQT QMRDGGLFSG GGR UniProtKB:  Tape measure protein Tape measure protein |
-Macromolecule #5: Tail tip assembly protein I
| Macromolecule | Name: Tail tip assembly protein I / type: protein_or_peptide / ID: 5 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia phage lambda (virus) Escherichia phage lambda (virus) |
| Molecular weight | Theoretical: 23.14659 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MAATHTLPLA SPGMARICLY GDLQRFGRRI DLRVKTGAEA IRALATQLPA FRQKLSDGWY QVRIAGRDVS TSGLTAQLHE TLPDGAVIH IVPRVAGAKS GGVFQIVLGA AAIAGSFFTA GATLAAWGAA IGAGGMTGIL FSLGASMVLG GVAQMLAPKA R TPRIQTTD ...String: MAATHTLPLA SPGMARICLY GDLQRFGRRI DLRVKTGAEA IRALATQLPA FRQKLSDGWY QVRIAGRDVS TSGLTAQLHE TLPDGAVIH IVPRVAGAKS GGVFQIVLGA AAIAGSFFTA GATLAAWGAA IGAGGMTGIL FSLGASMVLG GVAQMLAPKA R TPRIQTTD NGKQNTYFSS LDNMVAQGNV LPVLYGEMRV GSRVVSQEIS TADEGDGGQV VVIGR UniProtKB: Tail tip assembly protein I |
-Macromolecule #6: Tail tube protein
| Macromolecule | Name: Tail tube protein / type: protein_or_peptide / ID: 6 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Escherichia phage lambda (virus) Escherichia phage lambda (virus) |
| Molecular weight | Theoretical: 25.831779 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MPVPNPTMPV KGAGTTLWVY KGSGDPYANP LSDVDWSRLA KVKDLTPGEL TAESYDDSYL DDEDADWTAT GQGQKSAGDT SFTLAWMPG EQGQQALLAW FNEGDTRAYK IRFPNGTVDV FRGWVSSIGK AVTAKEVITR TVKVTNVGRP SMAEDRSTVT A ATGMTVTP ...String: MPVPNPTMPV KGAGTTLWVY KGSGDPYANP LSDVDWSRLA KVKDLTPGEL TAESYDDSYL DDEDADWTAT GQGQKSAGDT SFTLAWMPG EQGQQALLAW FNEGDTRAYK IRFPNGTVDV FRGWVSSIGK AVTAKEVITR TVKVTNVGRP SMAEDRSTVT A ATGMTVTP ASTSVVKGQS TTLTVAFQPE GVTDKSFRAV SADKTKATVS VSGMTITVNG VAAGKVNIPV VSGNGEFAAV AE ITVTAS UniProtKB: Tail tube protein |
-Macromolecule #7: IRON/SULFUR CLUSTER
| Macromolecule | Name: IRON/SULFUR CLUSTER / type: ligand / ID: 7 / Number of copies: 3 / Formula: SF4 |
|---|---|
| Molecular weight | Theoretical: 351.64 Da |
| Chemical component information |  ChemComp-FS1: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z
Z Y
Y X
X