[English] 日本語
 Yorodumi
Yorodumi- EMDB-34918: cellodextrin phosphorylase stable variant from Clostridium thermo... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cellodextrin phosphorylase stable variant from Clostridium thermocellum | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  cellulose / cellulose /  phosphorolysis / synthesis / phosphorolysis / synthesis /  CARBOHYDRATE CARBOHYDRATE | |||||||||
| Function / homology |  Function and homology information Function and homology information transferase activity / transferase activity /  carbohydrate binding / carbohydrate metabolic process carbohydrate binding / carbohydrate metabolic processSimilarity search - Function | |||||||||
| Biological species |   Acetivibrio thermocellus (bacteria) Acetivibrio thermocellus (bacteria) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.28 Å cryo EM / Resolution: 2.28 Å | |||||||||
 Authors Authors | Kuga T / Sunagawa N / Igarashi K | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: 11 cysteine-to-serine mutations improve the stability of cellodextrin phosphorylase Authors: Kuga T / Sunagawa N / Igarashi K | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34918.map.gz emd_34918.map.gz | 219.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34918-v30.xml emd-34918-v30.xml emd-34918.xml emd-34918.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
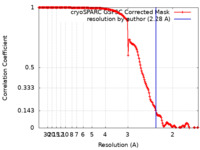
| FSC (resolution estimation) |  emd_34918_fsc.xml emd_34918_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_34918.png emd_34918.png | 139.4 KB | ||
| Filedesc metadata |  emd-34918.cif.gz emd-34918.cif.gz | 6.1 KB | ||
| Others |  emd_34918_half_map_1.map.gz emd_34918_half_map_1.map.gz emd_34918_half_map_2.map.gz emd_34918_half_map_2.map.gz | 391.3 MB 391.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34918 http://ftp.pdbj.org/pub/emdb/structures/EMD-34918 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34918 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34918 | HTTPS FTP |
-Related structure data
| Related structure data |  8hnuMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34918.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34918.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34918_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34918_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : cellodextrin phosphorylase dimer
| Entire | Name: cellodextrin phosphorylase dimer |
|---|---|
| Components |
|
-Supramolecule #1: cellodextrin phosphorylase dimer
| Supramolecule | Name: cellodextrin phosphorylase dimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Acetivibrio thermocellus (bacteria) / Strain: YM4 Acetivibrio thermocellus (bacteria) / Strain: YM4 |
| Molecular weight | Theoretical: 240 KDa |
-Macromolecule #1: Cellodextrin phosphorylase
| Macromolecule | Name: Cellodextrin phosphorylase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Acetivibrio thermocellus (bacteria) / Strain: YM4 Acetivibrio thermocellus (bacteria) / Strain: YM4 |
| Molecular weight | Theoretical: 112.736172 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: MGITKVTARN NKITPVELLN QKFGNKINLG NFADAVFTDA AFKNVAGIAN LPMKAPVMQV LMENSIVSKY LKQFVPDRSV SFVEEGQKF YIVLEDGQKI EVPEDVNKAL KATVSDVKHW AGYLTEDGEH VIDLLKPAPG PHFYVNLLIG NRLGFKRTLQ T TPKSVVDR ...String: MGITKVTARN NKITPVELLN QKFGNKINLG NFADAVFTDA AFKNVAGIAN LPMKAPVMQV LMENSIVSKY LKQFVPDRSV SFVEEGQKF YIVLEDGQKI EVPEDVNKAL KATVSDVKHW AGYLTEDGEH VIDLLKPAPG PHFYVNLLIG NRLGFKRTLQ T TPKSVVDR FGRGSFRSHA ATQVLATRFD MRQEENGFPA NRQFYLYEDG KQIFYSALID DNIVEATSKH SSNRTVIKYK TA SNLEITR TIFLVPHKKG FPLATELQRI EIKNASDKAR NLSITYTGMF GTGAVHAIFE DVTYTNVIMQ SAALYNDKGE FIG ITPDYY PEEFKQDTRF VTMIVRNGDE KSFPQSFSTD YNDFVGTGTL EHPAGGSNLN NKLNRKGPGF FALGAPFTVE PGKT VIIDT FTGLSSSKDN ENYSDAVMLR ELDNLLRYFE KSESVEETLN EIINFHENYG KYFQFNTGNK LFDSGFNRNL AFQVL YQTF MSRSFGQTQK GYREIGFREI QDLFASMYYF INIGYQDFVK ELLFEWTANV YKMGYANHNF YWVGKQPGLY SDDSLW LLQ AYYRYIIYTK DTSVLNEEVP VADGNNEKRA VRETLKAIIQ YSASISVGDH GLPLLDLADW NDSLKIDSNS IDGATKE KL YYEQLKKTNG KYGDRFMSDY SESVMNAFLL KLAIDHLAEI ATLDNDTQLA QQMSELSKEV TDRIQKHAWK ENFFARVL I NRYKDGSYTY LGAKGDKLSA DPNIDGVYFL NSFAWSVLSD VATDEQIAIM VDVIKKHLLT PYGLRLVTPA DLNKIANDT ATGHYFFGDR ENGAVFKHAS MMAVAALIKA AKKVKDNELA KEMARIAYFM IDLVLPYKNL ENPFQVAGNP RISTQYINTD TGENIGPLL SGTATWLNLN LISLAGIEYT RDGISFNPIL REEETQLNFT LKAPKSSYKF SITKPVGFAR MESSEYELFV D GQKIDNTV IPMYTDEKEH IVTLKFKLEH HHHHH UniProtKB:  Cellodextrin phosphorylase Cellodextrin phosphorylase |
-Macromolecule #2: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 316 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.0 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 49.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X