[English] 日本語
 Yorodumi
Yorodumi- EMDB-33003: Cryo-EM structure of Coxsackievirus B1 A-particle in complex with... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
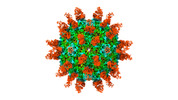
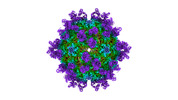
| Title | Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 9A3 (classified from CVB1 mature virion in complex with 8A10 and 9A3) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |   Coxsackievirus B1 Coxsackievirus B1 | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 4.01 Å cryo EM / Resolution: 4.01 Å | |||||||||
 Authors Authors | Zheng Q / Zhu R / Sun H / Cheng T / Li S / Xia N | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cell Host Microbe / Year: 2022 Journal: Cell Host Microbe / Year: 2022Title: Structural basis for the synergistic neutralization of coxsackievirus B1 by a triple-antibody cocktail. Authors: Qingbing Zheng / Rui Zhu / Zhichao Yin / Longfa Xu / Hui Sun / Hai Yu / Yuanyuan Wu / Yichao Jiang / Qiongzi Huang / Yang Huang / Dongqing Zhang / Liqin Liu / Hongwei Yang / Maozhou He / ...Authors: Qingbing Zheng / Rui Zhu / Zhichao Yin / Longfa Xu / Hui Sun / Hai Yu / Yuanyuan Wu / Yichao Jiang / Qiongzi Huang / Yang Huang / Dongqing Zhang / Liqin Liu / Hongwei Yang / Maozhou He / Zhenhong Zhou / Yanan Jiang / Zhenqin Chen / Huan Zhao / Yuqiong Que / Zhibo Kong / Lizhi Zhou / Tingting Li / Jun Zhang / Wenxin Luo / Ying Gu / Tong Cheng / Shaowei Li / Ningshao Xia /  Abstract: Coxsackievirus B1 (CVB1) is an emerging pathogen associated with severe neonatal diseases including aseptic meningitis, myocarditis, and pancreatitis and also with the development of type 1 diabetes. ...Coxsackievirus B1 (CVB1) is an emerging pathogen associated with severe neonatal diseases including aseptic meningitis, myocarditis, and pancreatitis and also with the development of type 1 diabetes. We characterize the binding and therapeutic efficacies of three CVB1-specific neutralizing antibodies (nAbs) identified for their ability to inhibit host receptor engagement. High-resolution cryo-EM structures showed that these antibodies recognize different epitopes but with an overlapping region in the capsid VP2 protein and specifically the highly variable EF loop. Moreover, they perturb capsid-receptor interactions by binding various viral particle forms. Antibody combinations achieve synergetic neutralization via a stepwise capsid transition and virion disruption, indicating dynamic changes in the virion in response to multiple nAbs targeting the receptor-binding site. Furthermore, this three-antibody cocktail protects against lethal challenge in neonatal mice and limits pancreatitis and viral replication in a non-obese diabetic mouse model. These results illustrate the utility of nAbs for rational design of therapeutics against picornaviruses such as CVB. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33003.map.gz emd_33003.map.gz | 632.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33003-v30.xml emd-33003-v30.xml emd-33003.xml emd-33003.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_33003.png emd_33003.png | 64.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33003 http://ftp.pdbj.org/pub/emdb/structures/EMD-33003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33003 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33003 | HTTPS FTP |
-Related structure data
| Related structure data |  7x2gC  7x2iC  7x2oC  7x2tC  7x2wC  7x37C  7x38C  7x3cC  7x3dC  7x3eC  7x3fC  7x40C  7x42C  7x46C  7x47C  7x49C  7x4kC  7x4mC C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33003.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33003.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Cryo-EM structure of Coxsackievirus B1 A-particle in complex with...
| Entire | Name: Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 9A3 (classified from CVB1 mature virion in complex with 8A10 and 9A3) |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of Coxsackievirus B1 A-particle in complex with...
| Supramolecule | Name: Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 9A3 (classified from CVB1 mature virion in complex with 8A10 and 9A3) type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:   Coxsackievirus B1 Coxsackievirus B1 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.9 µm / Nominal defocus min: 1.1 µm Bright-field microscopy / Nominal defocus max: 2.9 µm / Nominal defocus min: 1.1 µm |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
|---|---|
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.01 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1055 |
 Movie
Movie Controller
Controller