[English] 日本語
 Yorodumi
Yorodumi- EMDB-31428: Cryo-EM structure of the fibril formed by disaccharide-modified a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 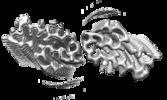 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the fibril formed by disaccharide-modified amyloid-beta(1-42) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationGolgi-associated vesicle /  clathrin-coated pit / clathrin-coated pit /  heparin binding / heparin binding /  growth cone / growth cone /  perikaryon / perikaryon /  early endosome / early endosome /  Golgi apparatus / Golgi apparatus /  cell surface / cell surface /  endoplasmic reticulum / extracellular region / endoplasmic reticulum / extracellular region /  nucleus nucleusSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
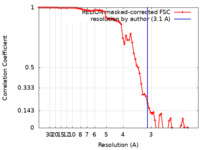
| Method | helical reconstruction /  cryo EM / Resolution: 3.1 Å cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Xia WC / Sun YP / Liu C | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: J.Am.Chem.Soc. / Year: 2021 Journal: J.Am.Chem.Soc. / Year: 2021Title: O-Glycosylation Induces Amyloid-beta To Form New Fibril Polymorphs Vulnerable for Degradation Authors: Liu D / Wei Q / Xia W / He C / Zhang Q / Huang L / Wang X / Sun Y / Ma Y / Zhang X / Wang Y / Shi X / Liu C / Dong S | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31428.map.gz emd_31428.map.gz | 4.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31428-v30.xml emd-31428-v30.xml emd-31428.xml emd-31428.xml | 9.4 KB 9.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_31428_fsc.xml emd_31428_fsc.xml | 7.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_31428.png emd_31428.png | 80.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31428 http://ftp.pdbj.org/pub/emdb/structures/EMD-31428 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31428 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31428 | HTTPS FTP |
-Related structure data
| Related structure data |  7f29MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31428.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31428.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Cryo-EM structure of the fibril formed by disaccharide-modified a...
| Entire | Name: Cryo-EM structure of the fibril formed by disaccharide-modified amyloid-beta(1-42) |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of the fibril formed by disaccharide-modified a...
| Supramolecule | Name: Cryo-EM structure of the fibril formed by disaccharide-modified amyloid-beta(1-42) type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Amyloid-beta A4 protein
| Macromolecule | Name: Amyloid-beta A4 protein / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.90044 KDa |
| Sequence | String: HDSGYEVHHQ KLVFFAEDVG SNKGAIIGLM VGGVVIA |
-Macromolecule #3: ACETIC ACID
| Macromolecule | Name: ACETIC ACID / type: ligand / ID: 3 / Number of copies: 6 / Formula: ACY |
|---|---|
| Molecular weight | Theoretical: 60.052 Da |
| Chemical component information |  ChemComp-ACY: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 55.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller