+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30665 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of histamine H1 receptor Gq complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information Histamine receptors / Histamine receptors /  histamine receptor activity / cellular response to histamine / histamine receptor activity / cellular response to histamine /  regulation of vascular permeability / G protein-coupled serotonin receptor activity / regulation of vascular permeability / G protein-coupled serotonin receptor activity /  neurotransmitter receptor activity / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / positive regulation of vasoconstriction / Olfactory Signaling Pathway / Activation of the phototransduction cascade ... neurotransmitter receptor activity / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / positive regulation of vasoconstriction / Olfactory Signaling Pathway / Activation of the phototransduction cascade ... Histamine receptors / Histamine receptors /  histamine receptor activity / cellular response to histamine / histamine receptor activity / cellular response to histamine /  regulation of vascular permeability / G protein-coupled serotonin receptor activity / regulation of vascular permeability / G protein-coupled serotonin receptor activity /  neurotransmitter receptor activity / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / positive regulation of vasoconstriction / Olfactory Signaling Pathway / Activation of the phototransduction cascade / neurotransmitter receptor activity / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / positive regulation of vasoconstriction / Olfactory Signaling Pathway / Activation of the phototransduction cascade /  regulation of synaptic plasticity / regulation of synaptic plasticity /  visual learning / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / visual learning / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G-protein activation / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / ADP signalling through P2Y purinoceptor 12 / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) /  memory / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / adenylate cyclase-activating dopamine receptor signaling pathway / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / G-protein beta-subunit binding / Inactivation, recovery and regulation of the phototransduction cascade / memory / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-type ligand receptors / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / cellular response to catecholamine stimulus / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / adenylate cyclase-activating dopamine receptor signaling pathway / G beta:gamma signalling through PI3Kgamma / cellular response to prostaglandin E stimulus / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / G-protein beta-subunit binding / Inactivation, recovery and regulation of the phototransduction cascade /  heterotrimeric G-protein complex / G alpha (12/13) signalling events / heterotrimeric G-protein complex / G alpha (12/13) signalling events /  extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / retina development in camera-type eye /  GTPase binding / phospholipase C-activating G protein-coupled receptor signaling pathway / Ca2+ pathway / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / G alpha (q) signalling events / chemical synaptic transmission / cell population proliferation / Ras protein signal transduction / Extra-nuclear estrogen signaling / GTPase binding / phospholipase C-activating G protein-coupled receptor signaling pathway / Ca2+ pathway / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / G alpha (q) signalling events / chemical synaptic transmission / cell population proliferation / Ras protein signal transduction / Extra-nuclear estrogen signaling /  inflammatory response / G protein-coupled receptor signaling pathway / lysosomal membrane / inflammatory response / G protein-coupled receptor signaling pathway / lysosomal membrane /  GTPase activity / GTPase activity /  synapse / synapse /  dendrite / protein-containing complex binding / dendrite / protein-containing complex binding /  signal transduction / extracellular exosome / signal transduction / extracellular exosome /  membrane / membrane /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
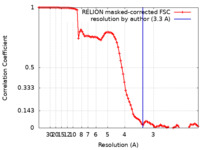
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.3 Å cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | He Y / Xia R / Wang N / Xu Z | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Cryo-EM structure of the human histamine H receptor/G complex. Authors: Ruixue Xia / Na Wang / Zhenmei Xu / Yang Lu / Jing Song / Anqi Zhang / Changyou Guo / Yuanzheng He /  Abstract: Histamine receptors play important roles in various pathophysiological conditions and are effective targets for anti-allergy treatment, however the mechanism of receptor activation remain elusive. ...Histamine receptors play important roles in various pathophysiological conditions and are effective targets for anti-allergy treatment, however the mechanism of receptor activation remain elusive. Here, we present the cryo-electron microscopy (cryo-EM) structure of the human HR in complex with a G protein in an active conformation via a NanoBiT tethering strategy. The structure reveals that histamine activates receptor via interacting with the key residues of both transmembrane domain 3 (TM3) and TM6 to squash the binding pocket on the extracellular side and to open the cavity on the intracellular side for G engagement in a model of "squash to activate and expand to deactivate". The structure also reveals features for G coupling, including the interaction between intracellular loop 2 (ICL2) and the αN-β junction of G protein. The detailed analysis of our structure will provide a framework for understanding G-protein coupling selectivity and clues for designing novel antihistamines. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30665.map.gz emd_30665.map.gz | 33 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30665-v30.xml emd-30665-v30.xml emd-30665.xml emd-30665.xml | 14.5 KB 14.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30665_fsc.xml emd_30665_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_30665.png emd_30665.png | 69 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30665 http://ftp.pdbj.org/pub/emdb/structures/EMD-30665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30665 | HTTPS FTP |
-Related structure data
| Related structure data |  7dflMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30665.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30665.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : GPCR/G-protein complex
| Entire | Name: GPCR/G-protein complex |
|---|---|
| Components |
|
-Supramolecule #1: GPCR/G-protein complex
| Supramolecule | Name: GPCR/G-protein complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Molecular weight | Theoretical: 130 kDa/nm |
-Macromolecule #1: Histamine H1 receptor
| Macromolecule | Name: Histamine H1 receptor / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.856035 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MSLPNSSCLL EDKMCEGNKT TMASPQLMPL VVVLSTICLV TVGLNLLVLY AVRSERKLHT VGNLYIVSLS VADLIVGAVV MPMNILYLL MSKWSLGRPL CLFWLSMDYV ASTASIFSVF ILCIDRYRSV QQPLRYLKYR TKTRASATIL GAWFLSFLWV I PILGWNHF ...String: MSLPNSSCLL EDKMCEGNKT TMASPQLMPL VVVLSTICLV TVGLNLLVLY AVRSERKLHT VGNLYIVSLS VADLIVGAVV MPMNILYLL MSKWSLGRPL CLFWLSMDYV ASTASIFSVF ILCIDRYRSV QQPLRYLKYR TKTRASATIL GAWFLSFLWV I PILGWNHF MQQTSVRRED KCETDFYDVT WFKVMTAIIN FYLPTLLMLW FYAKIYKAVR QHCQHRELIN RSLPSFSEIK LR PENPKGD AKKPGKESPW EVLKRKPKDA GGGSVLKSPS QTPKEMKSPV VFSQEDDREV DKLYCFPLDI VHMQAAAEGS SRD YVAVNR SHGQLKTDEQ GLNTHGASEI SEDQMLGDSQ SFSRTDSDTT TETAPGKGKL RSGSNTGLDY IKFTWKRLRS HSRQ YVSGL HMNRERKAAK QLGFIMAAFI LCWIPYFIFF MVIAFCKNCC NEHLHMFTIW LGYINSTLNP LIYPLCNENF KKTFK RILH IRS |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.285734 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW ...String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW DIETGQQTTT FTGHTGDVMS LSLAPDTRLF VSGACDASAK LWDVREGMCR QTFTGHESDI NAICFFPNGN AF ATGSDDA TCRLFDLRAD QELMTYSHDN IICGITSVSF SKSGRLLLAG YDDFNCNVWD ALKADRAGVL AGHDNRVSCL GVT DDGMAV ATGSWDSFLK IWN |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 6.375332 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: NTASIAQARK LVEQLKMEAN IDRIKVSKAA ADLMAYCEAH AKEDPLLTPV PASENPFR |
-Macromolecule #4: Guanine nucleotide-binding protein G(q) subunit alpha
| Macromolecule | Name: Guanine nucleotide-binding protein G(q) subunit alpha / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.090637 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: CTLSAEDKAA VERSKMIDRN LREDGEKARR ELKLLLLGTG ESGKSTFIKQ MRIIHGSGYS DEDKRGFTKL VYQNIFTAMQ AMIRAMDTL KIPYKHEHNK AHAQLVREVD VEKVSTFENP YVDAIRSLWS DPGIQECYDR RREYQLSDST KYYLNDLDRI A DSTYLPTQ ...String: CTLSAEDKAA VERSKMIDRN LREDGEKARR ELKLLLLGTG ESGKSTFIKQ MRIIHGSGYS DEDKRGFTKL VYQNIFTAMQ AMIRAMDTL KIPYKHEHNK AHAQLVREVD VEKVSTFENP YVDAIRSLWS DPGIQECYDR RREYQLSDST KYYLNDLDRI A DSTYLPTQ QDVLRVRVPT TGIIEYPFDL QSVIFRMVDV GGQRSERRKW IHCFENVTSI MFLVALSEYD QVLVESDNEN RM EESKALF RTIITYPWFQ NSSVILFLNK KDLLEEKIMY SHLVDYFPEY DGPQRDAQAA REFILKMFVD LNPDSDKILY SHF TCATDT ENIRFVFAAV KDTILQLNLK EYNLV |
-Macromolecule #5: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.679721 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAA |
-Macromolecule #6: HISTAMINE
| Macromolecule | Name: HISTAMINE / type: ligand / ID: 6 / Number of copies: 1 / Formula: HSM |
|---|---|
| Molecular weight | Theoretical: 111.145 Da |
| Chemical component information |  ChemComp-HSM: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 0.01 mm / Nominal defocus min: 1.2 µm Bright-field microscopy / Cs: 0.01 mm / Nominal defocus min: 1.2 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 56.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller