+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2936 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron cryo-microscopy structure of PB1-p62 filaments | |||||||||
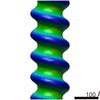
 Map data Map data | 3D reconstruction of PB1(1-102) p62 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Selective autophagy / autophagy receptor / autophagy scaffold / p62/SQSTM1 / single-particle helical reconstruction | |||||||||
| Function / homology |  Function and homology information Function and homology informationbrown fat cell proliferation / protein localization to perinuclear region of cytoplasm / response to stress /  protein binding / protein targeting to vacuole involved in autophagy / regulation of Ras protein signal transduction / protein binding / protein targeting to vacuole involved in autophagy / regulation of Ras protein signal transduction /  Lewy body / aggrephagy / response to mitochondrial depolarisation / positive regulation of mitophagy in response to mitochondrial depolarization ...brown fat cell proliferation / protein localization to perinuclear region of cytoplasm / response to stress / Lewy body / aggrephagy / response to mitochondrial depolarisation / positive regulation of mitophagy in response to mitochondrial depolarization ...brown fat cell proliferation / protein localization to perinuclear region of cytoplasm / response to stress /  protein binding / protein targeting to vacuole involved in autophagy / regulation of Ras protein signal transduction / protein binding / protein targeting to vacuole involved in autophagy / regulation of Ras protein signal transduction /  Lewy body / aggrephagy / response to mitochondrial depolarisation / positive regulation of mitophagy in response to mitochondrial depolarization / amphisome / negative regulation of toll-like receptor 4 signaling pathway / non-membrane-bounded organelle assembly / regulation of autophagy of mitochondrion / pexophagy / endosome organization / regulation of protein complex stability / protein heterooligomerization / autophagy of mitochondrion / molecular sequestering activity / phagophore assembly site / regulation of mitochondrion organization / Lewy body / aggrephagy / response to mitochondrial depolarisation / positive regulation of mitophagy in response to mitochondrial depolarization / amphisome / negative regulation of toll-like receptor 4 signaling pathway / non-membrane-bounded organelle assembly / regulation of autophagy of mitochondrion / pexophagy / endosome organization / regulation of protein complex stability / protein heterooligomerization / autophagy of mitochondrion / molecular sequestering activity / phagophore assembly site / regulation of mitochondrion organization /  aggresome / regulation of canonical NF-kappaB signal transduction / ubiquitin-modified protein reader activity / K63-linked polyubiquitin modification-dependent protein binding / Nuclear events mediated by NFE2L2 / aggresome / regulation of canonical NF-kappaB signal transduction / ubiquitin-modified protein reader activity / K63-linked polyubiquitin modification-dependent protein binding / Nuclear events mediated by NFE2L2 /  autolysosome / autolysosome /  temperature homeostasis / endosomal transport / temperature homeostasis / endosomal transport /  immune system process / intracellular non-membrane-bounded organelle / neurotrophin TRK receptor signaling pathway / immune system process / intracellular non-membrane-bounded organelle / neurotrophin TRK receptor signaling pathway /  mitophagy / positive regulation of macroautophagy / Signaling by ALK fusions and activated point mutants / mitophagy / positive regulation of macroautophagy / Signaling by ALK fusions and activated point mutants /  autophagosome / signaling adaptor activity / positive regulation of autophagy / autophagosome / signaling adaptor activity / positive regulation of autophagy /  energy homeostasis / energy homeostasis /  inclusion body / negative regulation of protein ubiquitination / sperm midpiece / inclusion body / negative regulation of protein ubiquitination / sperm midpiece /  ionotropic glutamate receptor binding / p75NTR recruits signalling complexes / PINK1-PRKN Mediated Mitophagy / Pexophagy / NRIF signals cell death from the nucleus / molecular condensate scaffold activity / NF-kB is activated and signals survival / ionotropic glutamate receptor binding / p75NTR recruits signalling complexes / PINK1-PRKN Mediated Mitophagy / Pexophagy / NRIF signals cell death from the nucleus / molecular condensate scaffold activity / NF-kB is activated and signals survival /  SH2 domain binding / SH2 domain binding /  sarcomere / protein sequestering activity / sarcomere / protein sequestering activity /  protein kinase C binding / protein kinase C binding /  ubiquitin binding / positive regulation of long-term synaptic potentiation / response to ischemia / ubiquitin binding / positive regulation of long-term synaptic potentiation / response to ischemia /  P-body / apoptotic signaling pathway / positive regulation of protein localization to plasma membrane / P-body / apoptotic signaling pathway / positive regulation of protein localization to plasma membrane /  macroautophagy / protein catabolic process / macroautophagy / protein catabolic process /  protein localization / PML body / protein localization / PML body /  receptor tyrosine kinase binding / receptor tyrosine kinase binding /  autophagy / Interleukin-1 signaling / protein import into nucleus / KEAP1-NFE2L2 pathway / protein-macromolecule adaptor activity / late endosome / autophagy / Interleukin-1 signaling / protein import into nucleus / KEAP1-NFE2L2 pathway / protein-macromolecule adaptor activity / late endosome /  signaling receptor activity / signaling receptor activity /  Neddylation / ubiquitin-dependent protein catabolic process / cytoplasmic vesicle / transcription by RNA polymerase II / Neddylation / ubiquitin-dependent protein catabolic process / cytoplasmic vesicle / transcription by RNA polymerase II /  cell differentiation / cell differentiation /  lysosome / lysosome /  endosome / intracellular signal transduction / positive regulation of protein phosphorylation / positive regulation of apoptotic process / endosome / intracellular signal transduction / positive regulation of protein phosphorylation / positive regulation of apoptotic process /  protein phosphorylation / intracellular membrane-bounded organelle / protein serine/threonine kinase activity / apoptotic process / protein phosphorylation / intracellular membrane-bounded organelle / protein serine/threonine kinase activity / apoptotic process /  ubiquitin protein ligase binding / protein-containing complex binding / negative regulation of apoptotic process / ubiquitin protein ligase binding / protein-containing complex binding / negative regulation of apoptotic process /  protein kinase binding / negative regulation of transcription by RNA polymerase II / protein kinase binding / negative regulation of transcription by RNA polymerase II /  enzyme binding / enzyme binding /  endoplasmic reticulum / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / endoplasmic reticulum / protein homodimerization activity / positive regulation of transcription by RNA polymerase II /  mitochondrion / extracellular exosome / zinc ion binding / mitochondrion / extracellular exosome / zinc ion binding /  nucleoplasm / identical protein binding nucleoplasm / identical protein bindingSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction /  cryo EM / Resolution: 10.9 Å cryo EM / Resolution: 10.9 Å | |||||||||
 Authors Authors | Ciuffa R / Lamark T / Tarafder A / Guesdon A / Rybina S / Hagen WJH / Johansen T / Sachse C | |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2015 Journal: Cell Rep / Year: 2015Title: The selective autophagy receptor p62 forms a flexible filamentous helical scaffold. Authors: Rodolfo Ciuffa / Trond Lamark / Abul K Tarafder / Audrey Guesdon / Sofia Rybina / Wim J H Hagen / Terje Johansen / Carsten Sachse /   Abstract: The scaffold protein p62/SQSTM1 is involved in protein turnover and signaling and is commonly found in dense protein bodies in eukaryotic cells. In autophagy, p62 acts as a selective autophagy ...The scaffold protein p62/SQSTM1 is involved in protein turnover and signaling and is commonly found in dense protein bodies in eukaryotic cells. In autophagy, p62 acts as a selective autophagy receptor that recognizes and shuttles ubiquitinated proteins to the autophagosome for degradation. The structural organization of p62 in cellular bodies and the interplay of these assemblies with ubiquitin and the autophagic marker LC3 remain to be elucidated. Here, we present a cryo-EM structural analysis of p62. Together with structures of assemblies from the PB1 domain, we show that p62 is organized in flexible polymers with the PB1 domain constituting a helical scaffold. Filamentous p62 is capable of binding LC3 and addition of long ubiquitin chains induces disassembly and shortening of filaments. These studies explain how p62 assemblies provide a large molecular scaffold for the nascent autophagosome and reveal how they can bind ubiquitinated cargo. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2936.map.gz emd_2936.map.gz | 2.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2936-v30.xml emd-2936-v30.xml emd-2936.xml emd-2936.xml | 23.3 KB 23.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2936.png emd_2936.png | 438.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2936 http://ftp.pdbj.org/pub/emdb/structures/EMD-2936 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2936 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2936 | HTTPS FTP |
-Related structure data
| Related structure data |  4uf8MC  2937C  4uf9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2936.map.gz / Format: CCP4 / Size: 4.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2936.map.gz / Format: CCP4 / Size: 4.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D reconstruction of PB1(1-102) p62 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : PB1(1-102) domain of p62/Sqstm1
| Entire | Name: PB1(1-102) domain of p62/Sqstm1 |
|---|---|
| Components |
|
-Supramolecule #1000: PB1(1-102) domain of p62/Sqstm1
| Supramolecule | Name: PB1(1-102) domain of p62/Sqstm1 / type: sample / ID: 1000 / Details: Helical polymer / Oligomeric state: Helical / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 11.3 KDa / Theoretical: 11.3 KDa / Method: Theoretical weight of construct |
-Macromolecule #1: Sequestosome-1
| Macromolecule | Name: Sequestosome-1 / type: protein_or_peptide / ID: 1 / Name.synonym: p62/SQSTM1 / Oligomeric state: Helical / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Experimental: 11.3 KDa / Theoretical: 11.3 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) / Recombinant strain: BL21 / Recombinant plasmid: pOPTM-p62-PB1_K103STOP_E104STOP Escherichia coli (E. coli) / Recombinant strain: BL21 / Recombinant plasmid: pOPTM-p62-PB1_K103STOP_E104STOP |
| Sequence | UniProtKB: Sequestosome-1 GO: phagophore assembly site, autophagy of mitochondrion, P-body, P-body, positive regulation of protein phosphorylation, immune system process, protein serine/threonine kinase activity, protein ...GO: phagophore assembly site, autophagy of mitochondrion,  P-body, P-body,  P-body, positive regulation of protein phosphorylation, P-body, positive regulation of protein phosphorylation,  immune system process, protein serine/threonine kinase activity, immune system process, protein serine/threonine kinase activity,  protein kinase C binding, protein kinase C binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  protein binding, protein binding,  nucleus, nucleus,  nucleus, nucleus,  nucleoplasm, nucleoplasm,  cytoplasm, cytoplasm,  cytoplasm, cytoplasm,  cytoplasm, cytoplasm,  lysosome, lysosome,  lysosome, lysosome,  endosome, late endosome, endosome, late endosome,  autophagosome, autophagosome,  autophagosome, autophagosome,  autophagosome, autophagosome,  endoplasmic reticulum, endoplasmic reticulum,  endoplasmic reticulum, endoplasmic reticulum,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  cytosol, cytosol,  protein phosphorylation, ubiquitin-dependent protein catabolic process, protein phosphorylation, ubiquitin-dependent protein catabolic process,  autophagy, autophagy,  autophagy, autophagy,  autophagy, apoptotic process, response to stress, autophagy, apoptotic process, response to stress,  protein localization, zinc ion binding, regulation of mitochondrion organization, endosomal transport, protein localization, zinc ion binding, regulation of mitochondrion organization, endosomal transport,  inclusion body, inclusion body,  aggresome, aggresome,  aggresome, aggresome,  macroautophagy, positive regulation of macroautophagy, PML body, macroautophagy, positive regulation of macroautophagy, PML body,  protein kinase binding, protein kinase binding,  cell differentiation, cell differentiation,  receptor tyrosine kinase binding, cytoplasmic vesicle, intracellular signal transduction, receptor tyrosine kinase binding, cytoplasmic vesicle, intracellular signal transduction,  SH2 domain binding, identical protein binding, identical protein binding, identical protein binding, identical protein binding, identical protein binding, protein homodimerization activity, positive regulation of apoptotic process, negative regulation of apoptotic process, regulation of canonical NF-kappaB signal transduction, SH2 domain binding, identical protein binding, identical protein binding, identical protein binding, identical protein binding, identical protein binding, protein homodimerization activity, positive regulation of apoptotic process, negative regulation of apoptotic process, regulation of canonical NF-kappaB signal transduction,  ubiquitin binding, positive regulation of transcription by RNA polymerase II, regulation of Ras protein signal transduction, ubiquitin binding, positive regulation of transcription by RNA polymerase II, regulation of Ras protein signal transduction,  metal ion binding, neurotrophin TRK receptor signaling pathway, neurotrophin TRK receptor signaling pathway, protein heterooligomerization, extracellular exosome, K63-linked polyubiquitin modification-dependent protein binding, apoptotic signaling pathway, positive regulation of mitophagy in response to mitochondrial depolarization, regulation of autophagy of mitochondrion metal ion binding, neurotrophin TRK receptor signaling pathway, neurotrophin TRK receptor signaling pathway, protein heterooligomerization, extracellular exosome, K63-linked polyubiquitin modification-dependent protein binding, apoptotic signaling pathway, positive regulation of mitophagy in response to mitochondrial depolarization, regulation of autophagy of mitochondrionInterPro: PB1 domain, UBA-like superfamily,  Ubiquitin-associated domain, Ubiquitin-associated domain,  Zinc finger, ZZ-type Zinc finger, ZZ-type |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 0.25 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 50 mM Tris pH 7.5, 100 mM NaCl, DTT 4 mM |
| Grid | Details: glow-discharged C-flat 1.2/1.3 and 200 mesh Quantifoil multi-A grids |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 77 K / Instrument: HOMEMADE PLUNGER / Method: Backside blotting |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 75000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Date | Feb 21, 2014 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Number real images: 994 / Average electron dose: 15 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: CTFFIND, convolution images, Wiener filter reconstruction |
|---|---|
| Final angle assignment | Details: SPIDER |
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 12.99 Å Applied symmetry - Helical parameters - Δ&Phi: 30.77 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 10.9 Å / Resolution method: OTHER / Software - Name: SPRING |
| Details | All of the image processing was carried using the SPRING package. |
 Movie
Movie Controller
Controller