[English] 日本語
 Yorodumi
Yorodumi- EMDB-26615: Complex of UBE2O with NAP1L1 and ubiquitylated uL2 (focused on UB... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Complex of UBE2O with NAP1L1 and ubiquitylated uL2 (focused on UBE2O-Ub) | ||||||||||||||||||
 Map data Map data | Sharpened map | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.5 Å cryo EM / Resolution: 3.5 Å | ||||||||||||||||||
 Authors Authors | Yip MCJ / Sedor SF / Shao S | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Mechanism of client selection by the protein quality-control factor UBE2O. Authors: Matthew C J Yip / Samantha F Sedor / Sichen Shao /  Abstract: The E2/E3 enzyme UBE2O ubiquitylates diverse clients to mediate important processes, including targeting unassembled 'orphan' proteins for quality control and clearing ribosomes during erythropoiesis. ...The E2/E3 enzyme UBE2O ubiquitylates diverse clients to mediate important processes, including targeting unassembled 'orphan' proteins for quality control and clearing ribosomes during erythropoiesis. How quality-control factors, such as UBE2O, select clients on the basis of heterogeneous features is largely unknown. Here, we show that UBE2O client selection is regulated by ubiquitin binding and a cofactor, NAP1L1. Attaching a single ubiquitin onto a client enhances UBE2O binding and multi-mono-ubiquitylation. UBE2O also repurposes the histone chaperone NAP1L1 as an adapter to recruit a subset of clients. Cryo-EM structures of human UBE2O in complex with NAP1L1 reveal a malleable client recruitment interface that is autoinhibited by the intrinsically reactive UBC domain. Adding a ubiquitylated client identifies a distinct ubiquitin-binding SH3-like domain required for client selection. Our findings reveal how multivalency and a feed-forward mechanism drive the selection of protein quality-control clients. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26615.map.gz emd_26615.map.gz | 13.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26615-v30.xml emd-26615-v30.xml emd-26615.xml emd-26615.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26615_fsc.xml emd_26615_fsc.xml | 12.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_26615.png emd_26615.png | 77.6 KB | ||
| Others |  emd_26615_additional_1.map.gz emd_26615_additional_1.map.gz emd_26615_additional_2.map.gz emd_26615_additional_2.map.gz emd_26615_half_map_1.map.gz emd_26615_half_map_1.map.gz emd_26615_half_map_2.map.gz emd_26615_half_map_2.map.gz | 133 MB 117.1 MB 118.3 MB 118.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26615 http://ftp.pdbj.org/pub/emdb/structures/EMD-26615 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26615 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26615 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26615.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26615.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: deepEMhancer sharpened map
| File | emd_26615_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | deepEMhancer sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
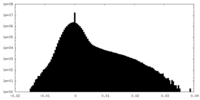
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_26615_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_26615_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
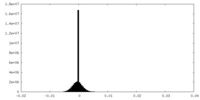
| Density Histograms |
-Half map: Half map 2
| File | emd_26615_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of UBE2O with NAP1L1 and ubiquitylated uL2 (focused on UB...
| Entire | Name: Complex of UBE2O with NAP1L1 and ubiquitylated uL2 (focused on UBE2O-Ub) |
|---|---|
| Components |
|
-Supramolecule #1: Complex of UBE2O with NAP1L1 and ubiquitylated uL2 (focused on UB...
| Supramolecule | Name: Complex of UBE2O with NAP1L1 and ubiquitylated uL2 (focused on UBE2O-Ub) type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.4000000000000001 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.4000000000000001 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 59.6 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)