[English] 日本語
 Yorodumi
Yorodumi- EMDB-26015: Complex GGNN of AMPA-subtype iGluR GluA2 in complex with auxiliar... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Complex GGNN of AMPA-subtype iGluR GluA2 in complex with auxiliary subunit gamma2 (Stargazin) at low glutamate concentration (20 uM) in the presence of cyclothiazide (100 uM) | |||||||||||||||
 Map data Map data | Complex GGNN of AMPA-subtype iGluR GluA2 in complex with auxiliary subunit gamma2 (Stargazin) at low glutamate concentration (20 uM) in the presence of cyclothiazide (100 uM) | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationspine synapse / dendritic spine neck / dendritic spine head / Activation of AMPA receptors / response to lithium ion / perisynaptic space / cellular response to glycine / AMPA glutamate receptor activity / Trafficking of GluR2-containing AMPA receptors /  immunoglobulin binding ...spine synapse / dendritic spine neck / dendritic spine head / Activation of AMPA receptors / response to lithium ion / perisynaptic space / cellular response to glycine / AMPA glutamate receptor activity / Trafficking of GluR2-containing AMPA receptors / immunoglobulin binding ...spine synapse / dendritic spine neck / dendritic spine head / Activation of AMPA receptors / response to lithium ion / perisynaptic space / cellular response to glycine / AMPA glutamate receptor activity / Trafficking of GluR2-containing AMPA receptors /  immunoglobulin binding / AMPA glutamate receptor complex / kainate selective glutamate receptor activity / immunoglobulin binding / AMPA glutamate receptor complex / kainate selective glutamate receptor activity /  ionotropic glutamate receptor complex / extracellularly glutamate-gated ion channel activity / asymmetric synapse / regulation of receptor recycling / Unblocking of NMDA receptors, glutamate binding and activation / ionotropic glutamate receptor complex / extracellularly glutamate-gated ion channel activity / asymmetric synapse / regulation of receptor recycling / Unblocking of NMDA receptors, glutamate binding and activation /  glutamate receptor binding / positive regulation of synaptic transmission / glutamate-gated receptor activity / presynaptic active zone membrane / response to fungicide / cellular response to brain-derived neurotrophic factor stimulus / glutamate receptor binding / positive regulation of synaptic transmission / glutamate-gated receptor activity / presynaptic active zone membrane / response to fungicide / cellular response to brain-derived neurotrophic factor stimulus /  regulation of synaptic transmission, glutamatergic / somatodendritic compartment / dendrite membrane / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / regulation of synaptic transmission, glutamatergic / somatodendritic compartment / dendrite membrane / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential /  ionotropic glutamate receptor binding / ionotropic glutamate receptor signaling pathway / dendrite cytoplasm / ionotropic glutamate receptor binding / ionotropic glutamate receptor signaling pathway / dendrite cytoplasm /  cytoskeletal protein binding / cytoskeletal protein binding /  SNARE binding / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / dendritic shaft / SNARE binding / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / dendritic shaft /  synaptic membrane / synaptic membrane /  synaptic transmission, glutamatergic / synaptic transmission, glutamatergic /  PDZ domain binding / postsynaptic density membrane / protein tetramerization / modulation of chemical synaptic transmission / Schaffer collateral - CA1 synapse / establishment of protein localization / PDZ domain binding / postsynaptic density membrane / protein tetramerization / modulation of chemical synaptic transmission / Schaffer collateral - CA1 synapse / establishment of protein localization /  terminal bouton / terminal bouton /  receptor internalization / synaptic vesicle membrane / cerebral cortex development / receptor internalization / synaptic vesicle membrane / cerebral cortex development /  synaptic vesicle / presynapse / synaptic vesicle / presynapse /  presynaptic membrane / presynaptic membrane /  signaling receptor activity / signaling receptor activity /  amyloid-beta binding / amyloid-beta binding /  growth cone / growth cone /  perikaryon / chemical synaptic transmission / perikaryon / chemical synaptic transmission /  scaffold protein binding / scaffold protein binding /  postsynaptic membrane / postsynaptic membrane /  dendritic spine / dendritic spine /  postsynaptic density / neuron projection / postsynaptic density / neuron projection /  axon / neuronal cell body / axon / neuronal cell body /  dendrite / dendrite /  synapse / glutamatergic synapse / protein-containing complex binding / endoplasmic reticulum membrane / synapse / glutamatergic synapse / protein-containing complex binding / endoplasmic reticulum membrane /  protein kinase binding / protein kinase binding /  cell surface / cell surface /  endoplasmic reticulum / protein-containing complex / endoplasmic reticulum / protein-containing complex /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||||||||
| Biological species |   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) | |||||||||||||||
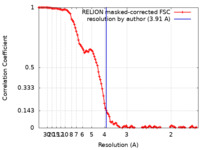
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.91 Å cryo EM / Resolution: 3.91 Å | |||||||||||||||
 Authors Authors | Yelshanskaya MV / Sobolevsky AI | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Opening of glutamate receptor channel to subconductance levels. Authors: Maria V Yelshanskaya / Dhilon S Patel / Christopher M Kottke / Maria G Kurnikova / Alexander I Sobolevsky /  Abstract: Ionotropic glutamate receptors (iGluRs) are tetrameric ligand-gated ion channels that open their pores in response to binding of the agonist glutamate. An ionic current through a single iGluR channel ...Ionotropic glutamate receptors (iGluRs) are tetrameric ligand-gated ion channels that open their pores in response to binding of the agonist glutamate. An ionic current through a single iGluR channel shows up to four discrete conductance levels (O1-O4). Higher conductance levels have been associated with an increased number of agonist molecules bound to four individual ligand-binding domains (LBDs). Here we determine structures of a synaptic complex of AMPA-subtype iGluR and the auxiliary subunit γ2 in non-desensitizing conditions with various occupancy of the LBDs by glutamate. We show that glutamate binds to LBDs of subunits B and D only after it is already bound to at least the same number of LBDs that belong to subunits A and C. Our structures combined with single-channel recordings, molecular dynamics simulations and machine-learning analysis suggest that channel opening requires agonist binding to at least two LBDs. Conversely, agonist binding to all four LBDs does not guarantee maximal channel conductance and favours subconductance states O1 and O2, with O3 and O4 being rare and not captured structurally. The lack of subunit independence and low efficiency coupling of glutamate binding to channel opening underlie the gating of synaptic complexes to submaximal conductance levels, which provide a potential for upregulation of synaptic activity. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26015.map.gz emd_26015.map.gz | 11.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26015-v30.xml emd-26015-v30.xml emd-26015.xml emd-26015.xml | 13.7 KB 13.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26015_fsc.xml emd_26015_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_26015.png emd_26015.png | 60.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26015 http://ftp.pdbj.org/pub/emdb/structures/EMD-26015 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26015 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26015 | HTTPS FTP |
-Related structure data
| Related structure data |  7tnnMC  7tnjC  7tnkC  7tnlC  7tnmC  7tnoC  7tnpC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26015.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26015.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Complex GGNN of AMPA-subtype iGluR GluA2 in complex with auxiliary subunit gamma2 (Stargazin) at low glutamate concentration (20 uM) in the presence of cyclothiazide (100 uM) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Ionotropic glutamate receptor GluA2 in complex with auxiliary sub...
| Entire | Name: Ionotropic glutamate receptor GluA2 in complex with auxiliary subunit gamma2 or Stargazin |
|---|---|
| Components |
|
-Supramolecule #1: Ionotropic glutamate receptor GluA2 in complex with auxiliary sub...
| Supramolecule | Name: Ionotropic glutamate receptor GluA2 in complex with auxiliary subunit gamma2 or Stargazin type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Isoform Flip of Glutamate receptor 2,Voltage-dependent calcium ch...
| Macromolecule | Name: Isoform Flip of Glutamate receptor 2,Voltage-dependent calcium channel gamma-3 subunit chimera type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) |
| Molecular weight | Theoretical: 115.444922 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NSIQIGGLFP RGADQEYSAF RVGMVQFSTS EFRLTPHIDN LEVANSFAVT NAFCSQFSRG VYAIFGFYDK KSVNTITSFC GTLHVSFIT PSFPTDGTHP FVIQMRPDLK GALLSLIEYY QWDKFAYLYD SDRGLSTLQA VLDSAAEKKW QVTAINVGNI N NDKKDETY ...String: NSIQIGGLFP RGADQEYSAF RVGMVQFSTS EFRLTPHIDN LEVANSFAVT NAFCSQFSRG VYAIFGFYDK KSVNTITSFC GTLHVSFIT PSFPTDGTHP FVIQMRPDLK GALLSLIEYY QWDKFAYLYD SDRGLSTLQA VLDSAAEKKW QVTAINVGNI N NDKKDETY RSLFQDLELK KERRVILDCE RDKVNDIVDQ VITIGKHVKG YHYIIANLGF TDGDLLKIQF GGAEVSGFQI VD YDDSLVS KFIERWSTLE EKEYPGAHTA TIKYTSALTY DAVQVMTEAF RNLRKQRIEI SRRGNAGDCL ANPAVPWGQG VEI ERALKQ VQVEGLSGNI KFDQNGKRIN YTINIMELKT NGPRKIGYWS EVDKMVLTED DTSGLEQKTV VVTTILESPY VMMK KNHEM LEGNERYEGY CVDLAAEIAK HCGFKYKLTI VGDGKYGARD ADTKIWNGMV GELVYGKADI AIAPLTITLV REEVI DFSK PFMSLGISIM IKKPQKSKPG VFSFLDPLAY EIWMCIVFAY IGVSVVLFLV SRFSPYEWHT EEFEDGRETQ SSESTN EFG IFNSLWFSLG AFMQQGCDIS PRSLSGRIVG GVWWFFTLII ISSYTANLAA FLTVERMVSP IESAEDLSKQ TEIAYGT LD SGSTKEFFRR SKIAVFDKMW TYMRSAEPSV FVRTTAEGVA RVRKSKGKYA YLLESTMNEY IEQRKPCDTM KVGGNLDS K GYGIATPKGS SLGTPVNLAV LKLSEQGVLD KLKNKWWYDK GECGAKDSGS KEKTSALSLS NVAGVFYILV GGLGLAMLV ALIEFCYKSR AEAKRMKGTG LFDRGVQMLL TTVGAFAAFS LMTIAVGTDY WLYSRGVCKT KSVSEDETSK KNEEVMTHSG LWRTCCLEG NFKGLCKQID HFPEDADYEA DTAEYFLRAV RASSIFPILS VILLFMGGLC IAASEFYKTR HNIILSAGIF F VSAGLSNI IGIIVYISAN AGDPSKSDSK KNSYSYGWSF YFGALSFIIA EMVGVLAVHM FIDRHKQLTG GLVPR |
-Macromolecule #2: GLUTAMIC ACID
| Macromolecule | Name: GLUTAMIC ACID / type: ligand / ID: 2 / Number of copies: 2 / Formula: GLU |
|---|---|
| Molecular weight | Theoretical: 147.129 Da |
| Chemical component information |  ChemComp-GLU: |
-Macromolecule #3: CYCLOTHIAZIDE
| Macromolecule | Name: CYCLOTHIAZIDE / type: ligand / ID: 3 / Number of copies: 4 / Formula: CYZ |
|---|---|
| Molecular weight | Theoretical: 389.878 Da |
| Chemical component information |  ChemComp-CYZ: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 58.5 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)