[English] 日本語
 Yorodumi
Yorodumi- EMDB-25609: E. coli DnaB bound to three DnaG C-terminal domains, ssDNA, ADP a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 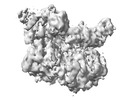 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli DnaB bound to three DnaG C-terminal domains, ssDNA, ADP and AlF4 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  helicase / SF4 / helicase / SF4 /  AAA+ / REPLICATION-DNA complex AAA+ / REPLICATION-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationDnaB-DnaC complex / DnaB-DnaC-Rep-PriC complex / DnaB-DnaG complex / DnaB-DnaC-DnaT-PriA-PriC complex / DNA primase DnaG /  DNA helicase complex / DnaB-DnaC-DnaT-PriA-PriB complex / DNA helicase complex / DnaB-DnaC-DnaT-PriA-PriB complex /  primosome complex / primosome complex /  DNA primase activity / DNA primase activity /  DNA replication, synthesis of primer ...DnaB-DnaC complex / DnaB-DnaC-Rep-PriC complex / DnaB-DnaG complex / DnaB-DnaC-DnaT-PriA-PriC complex / DNA primase DnaG / DNA replication, synthesis of primer ...DnaB-DnaC complex / DnaB-DnaC-Rep-PriC complex / DnaB-DnaG complex / DnaB-DnaC-DnaT-PriA-PriC complex / DNA primase DnaG /  DNA helicase complex / DnaB-DnaC-DnaT-PriA-PriB complex / DNA helicase complex / DnaB-DnaC-DnaT-PriA-PriB complex /  primosome complex / primosome complex /  DNA primase activity / DNA primase activity /  DNA replication, synthesis of primer / DNA replication, synthesis of primer /  replisome / DNA duplex unwinding / response to ionizing radiation / DNA unwinding involved in DNA replication / replication fork processing / DNA replication initiation / replisome / DNA duplex unwinding / response to ionizing radiation / DNA unwinding involved in DNA replication / replication fork processing / DNA replication initiation /  DNA helicase activity / DNA helicase activity /  DNA-directed RNA polymerase complex / DNA-directed RNA polymerase complex /  helicase activity / helicase activity /  DNA helicase / DNA helicase /  DNA replication / DNA replication /  ATP hydrolysis activity / ATP hydrolysis activity /  DNA binding / zinc ion binding / DNA binding / zinc ion binding /  ATP binding / identical protein binding / ATP binding / identical protein binding /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Escherichia coli K-12 (bacteria) / Escherichia coli K-12 (bacteria) /   Escherichia coli (E. coli) Escherichia coli (E. coli) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 4.2 Å cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Oakley AJ / Xu ZQ | |||||||||
| Funding support |  Australia, 1 items Australia, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Sequence of conformation changes of DnaB helicase during DNA unwinding and priming in Escherichia coli Authors: Oakley AJ / Xu ZQ | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25609.map.gz emd_25609.map.gz | 65.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25609-v30.xml emd-25609-v30.xml emd-25609.xml emd-25609.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25609.png emd_25609.png | 68.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25609 http://ftp.pdbj.org/pub/emdb/structures/EMD-25609 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25609 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25609 | HTTPS FTP |
-Related structure data
| Related structure data |  7t22MC  7t20C  7t21C  7t23C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25609.map.gz / Format: CCP4 / Size: 70.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25609.map.gz / Format: CCP4 / Size: 70.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.94 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : E. coli DnaB DnaG C-terminal domain bound to ssDNA
| Entire | Name: E. coli DnaB DnaG C-terminal domain bound to ssDNA |
|---|---|
| Components |
|
-Supramolecule #1: E. coli DnaB DnaG C-terminal domain bound to ssDNA
| Supramolecule | Name: E. coli DnaB DnaG C-terminal domain bound to ssDNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:   Escherichia coli K-12 (bacteria) Escherichia coli K-12 (bacteria) |
-Macromolecule #1: Replicative DNA helicase
| Macromolecule | Name: Replicative DNA helicase / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO / EC number:  DNA helicase DNA helicase |
|---|---|
| Source (natural) | Organism:   Escherichia coli K-12 (bacteria) / Strain: K12 Escherichia coli K-12 (bacteria) / Strain: K12 |
| Molecular weight | Theoretical: 52.406918 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: MAGNKPFNKQ QAEPRERDPQ VAGLKVPPHS IEAEQSVLGG LMLDNERWDD VAERVVADDF YTRPHRHIFT EMARLQESGS PIDLITLAE SLERQGQLDS VGGCAYLAEL SKNTPSAANI SAYADIVRER AVVREMISVA NEIAEAGFDP QGRTSEDLLD L AESRVFKI ...String: MAGNKPFNKQ QAEPRERDPQ VAGLKVPPHS IEAEQSVLGG LMLDNERWDD VAERVVADDF YTRPHRHIFT EMARLQESGS PIDLITLAE SLERQGQLDS VGGCAYLAEL SKNTPSAANI SAYADIVRER AVVREMISVA NEIAEAGFDP QGRTSEDLLD L AESRVFKI AESRANKDEG PKNIADVLDA TVARIEQLFQ QPHDGVTGVN TGYDDLNKKT AGLQPSDLII VAARPSMGKT TF AMNLVEN AAMLQDKPVL IFSLEMPSEQ IMMRSLASLS RVDQTKIRTG QLDDEDWARI SGTMGILLEK RNIYIDDSSG LTP TEVRSR ARRIAREHGG IGLIMIDYLQ LMRVPALSDN RTLEIAEISR SLKALAKELN VPVVALSQLN RSLEQRADKR PVNS DLRES GSIEQDADLI MFIYRDEVYH ENSDLKGIAE IIIGKQRNGP IGTVRLTFNG QWSRFDNYAG PQYDDE |
-Macromolecule #2: DNA primase
| Macromolecule | Name: DNA primase / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: DNA primase DnaG |
|---|---|
| Source (natural) | Organism:   Escherichia coli K-12 (bacteria) / Strain: K12 Escherichia coli K-12 (bacteria) / Strain: K12 |
| Molecular weight | Theoretical: 16.664918 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: AAESGVSRPV PQLKRTTMRI LIGLLVQNPE LATLVPPLEN LDENKLPGLG LFRELVNTCL SQPGLTTGQL LEHYRGTNNA ATLEKLSMW DDIADKNIAE QTFTDSLNHM FDSLLELRQE ELIARERTHG LSNEECLELW TLNQELAKK |
-Macromolecule #3: DNA (5'-D(*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*...
| Macromolecule | Name: DNA (5'-D(*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*TP*T)-3') type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Molecular weight | Theoretical: 6.038899 KDa |
| Sequence | String: (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) |
-Macromolecule #4: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 5 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #5: TETRAFLUOROALUMINATE ION
| Macromolecule | Name: TETRAFLUOROALUMINATE ION / type: ligand / ID: 5 / Number of copies: 5 / Formula: ALF |
|---|---|
| Molecular weight | Theoretical: 102.975 Da |
| Chemical component information |  ChemComp-ALF: |
-Macromolecule #6: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 6 / Number of copies: 5 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.6 Details: 20 mM Tris-HCl, pH 7.6, 100 mM NaCl, 5 mM MgCl2, 3 mM DTT, 0.25 mM EDTA and 100 micromolar ADP. |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE Details: Grids covered with a 2 nm continuous carbon film. Glow-discharged grids were floated on 40 microL drops of a 0.1% (w/v) aqueous solution of poly-lysine hydrobromide (MW 30-70 kDa; ...Details: Grids covered with a 2 nm continuous carbon film. Glow-discharged grids were floated on 40 microL drops of a 0.1% (w/v) aqueous solution of poly-lysine hydrobromide (MW 30-70 kDa; Polysciences, Inc.) for two min, briefly rinsed with MilliQ water 4 times and then allow to air dry. |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV Details: 3 microL of sample was applied to glow-discharged grids. Grids were blotted at 6 degrees C for 3.5 s with no extra blot force.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 657240 |
|---|---|
| Startup model | Type of model: NONE |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.07) |
| Final 3D classification | Number classes: 2 / Software - Name: RELION (ver. 3.07) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.07) |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 4.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.07) / Number images used: 46096 |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Target criteria: Correlation Coefficient |
|---|---|
| Output model |  PDB-7t22: |
 Movie
Movie Controller
Controller