+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2432 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of the COPII coat assembled on membranes | |||||||||
 Map data Map data | pos277.mrc | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | COPII / coat / secretion / trafficking / Sec23 / Sec24 / Sar1 / Sec13 / Sec31 / membrane / budding | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of COPII vesicle coating / COPII-coated vesicle cargo loading / vesicle organization / COPII vesicle coat / positive regulation of protein exit from endoplasmic reticulum / membrane organization / endoplasmic reticulum exit site / endoplasmic reticulum to Golgi vesicle-mediated transport / SNARE binding / intracellular protein transport ...regulation of COPII vesicle coating / COPII-coated vesicle cargo loading / vesicle organization / COPII vesicle coat / positive regulation of protein exit from endoplasmic reticulum / membrane organization / endoplasmic reticulum exit site / endoplasmic reticulum to Golgi vesicle-mediated transport / SNARE binding / intracellular protein transport / Golgi membrane / GTPase activity / endoplasmic reticulum membrane / GTP binding / Golgi apparatus / zinc ion binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM / negative staining | |||||||||
 Authors Authors | Zanetti G / Prinz S / Daum S / Meister A / Schekman R / Bacia K / Briggs JAG | |||||||||
 Citation Citation |  Journal: Elife / Year: 2013 Journal: Elife / Year: 2013Title: The structure of the COPII transport-vesicle coat assembled on membranes. Authors: Giulia Zanetti / Simone Prinz / Sebastian Daum / Annette Meister / Randy Schekman / Kirsten Bacia / John A G Briggs /  Abstract: Coat protein complex II (COPII) mediates formation of the membrane vesicles that export newly synthesised proteins from the endoplasmic reticulum. The inner COPII proteins bind to cargo and membrane, ...Coat protein complex II (COPII) mediates formation of the membrane vesicles that export newly synthesised proteins from the endoplasmic reticulum. The inner COPII proteins bind to cargo and membrane, linking them to the outer COPII components that form a cage around the vesicle. Regulated flexibility in coat architecture is essential for transport of a variety of differently sized cargoes, but structural data on the assembled coat has not been available. We have used cryo-electron tomography and subtomogram averaging to determine the structure of the complete, membrane-assembled COPII coat. We describe a novel arrangement of the outer coat and find that the inner coat can assemble into regular lattices. The data reveal how coat subunits interact with one another and with the membrane, suggesting how coordinated assembly of inner and outer coats can mediate and regulate packaging of vesicles ranging from small spheres to large tubular carriers. DOI:http://dx.doi.org/10.7554/eLife.00951.001. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2432.map.gz emd_2432.map.gz | 4.1 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2432-v30.xml emd-2432-v30.xml emd-2432.xml emd-2432.xml | 16.8 KB 16.8 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2432.png EMD-2432.png | 389.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2432 http://ftp.pdbj.org/pub/emdb/structures/EMD-2432 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2432 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2432 | HTTPS FTP |
-Validation report
| Summary document |  emd_2432_validation.pdf.gz emd_2432_validation.pdf.gz | 137.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2432_full_validation.pdf.gz emd_2432_full_validation.pdf.gz | 137.1 KB | Display | |
| Data in XML |  emd_2432_validation.xml.gz emd_2432_validation.xml.gz | 4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2432 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2432 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2432 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2432 | HTTPS FTP |
-Related structure data
| Related structure data |  2428C  2429C  2430C 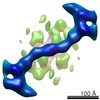 2431C  4bziC  4bzjC  4bzkC C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2432.map.gz / Format: CCP4 / Size: 4.6 GB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_2432.map.gz / Format: CCP4 / Size: 4.6 GB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | pos277.mrc | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Cryo-tomogram of COPII-coated membrane
| Entire | Name: Cryo-tomogram of COPII-coated membrane |
|---|---|
| Components |
|
-Supramolecule #1000: Cryo-tomogram of COPII-coated membrane
| Supramolecule | Name: Cryo-tomogram of COPII-coated membrane / type: sample / ID: 1000 / Number unique components: 5 |
|---|
-Macromolecule #1: Sar1p
| Macromolecule | Name: Sar1p / type: protein_or_peptide / ID: 1 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 21.437 KDa / Theoretical: 21.437 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Small COPII coat GTPase SAR1 / InterPro: Ras GTPase-activating domain, Roc domain |
-Macromolecule #2: Sec23p
| Macromolecule | Name: Sec23p / type: protein_or_peptide / ID: 2 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 85.437 KDa / Theoretical: 85.437 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: UNIPROTKB: E7QAP0 InterPro: Sec23/Sec24, trunk domain, Sec23/Sec24, helical domain, Sec23/Sec24 beta-sandwich, Zinc finger, Sec23/Sec24-type |
-Macromolecule #3: Sec24p
| Macromolecule | Name: Sec24p / type: protein_or_peptide / ID: 3 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 103.577 KDa / Theoretical: 103.577 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Sec24p InterPro: Sec23/Sec24, trunk domain, Sec23/Sec24 beta-sandwich, Sec23/Sec24, helical domain, Zinc finger, Sec23/Sec24-type |
-Macromolecule #4: Sec13p
| Macromolecule | Name: Sec13p / type: protein_or_peptide / ID: 4 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 20.794 KDa / Theoretical: 20.794 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: UNIPROTKB: E7Q6Z3 |
-Macromolecule #5: Sec31p
| Macromolecule | Name: Sec31p / type: protein_or_peptide / ID: 5 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 138.824 KDa / Theoretical: 138.824 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: UNIPROTKB: E7Q1I6 |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 / Details: HEPES, 50 mM KOAc, 1.2 mM MgCl2 |
|---|---|
| Staining | Type: NEGATIVE / Details: plunge frozen |
| Grid | Details: C-flat grids |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
- Electron microscopy #1
Electron microscopy #1
| Microscopy ID | 1 |
|---|---|
| Microscope | FEI TITAN KRIOS |
| Specialist optics | Energy filter - Name: GATAN GIF 2002 |
| Date | Sep 18, 2012 |
| Image recording | Category: CCD / Film or detector model: GATAN MULTISCAN / Number real images: 26 / Average electron dose: 80 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.2 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 19500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Tilt series - Axis1 - Min angle: -60 ° / Tilt series - Axis1 - Max angle: 60 ° / Tilt series - Axis1 - Angle increment: 3 ° |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Electron microscopy #2
Electron microscopy #2
| Microscopy ID | 2 |
|---|---|
| Microscope | FEI TITAN KRIOS |
| Specialist optics | Energy filter - Name: GATAN GIF 2002 |
| Date | Jun 19, 2012 |
| Image recording | Category: CCD / Film or detector model: GATAN MULTISCAN / Number real images: 26 / Average electron dose: 80 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.2 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 19500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Tilt series - Axis1 - Min angle: -60 ° / Tilt series - Axis1 - Max angle: 60 ° / Tilt series - Axis1 - Angle increment: 3 ° |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | CTF correction was performed on each projection. Projections aligned based on gold fiducially |
|---|---|
| Final reconstruction | Software - Name: Imod, Raptor / Number images used: 40 |
| CTF correction | Details: each projection |
 Movie
Movie Controller
Controller








