[English] 日本語
 Yorodumi
Yorodumi- EMDB-18808: SD1-3 Fab in complex with SARS-CoV-2 BA.2.12.1 Spike Glycoprotein -
+ Open data
Open data
- Basic information
Basic information
| Entry | 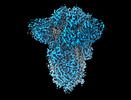 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | SD1-3 Fab in complex with SARS-CoV-2 BA.2.12.1 Spike Glycoprotein | |||||||||||||||
 Map data Map data | CryoEM sharpened C3 symmetric map of SD1-6 fab in complex with BA.2.12.1 S. | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords |  SARS-CoV-2 / Spike / SARS-CoV-2 / Spike /  Glycoprotein / Glycoprotein /  Coronavirus / Coronavirus /  antibody / antibody /  Fab / SD1 domain / Fab / SD1 domain /  therapeutic / therapeutic /  complex / neutralisingm convalescent sera / complex / neutralisingm convalescent sera /  viral protein / viral protein /  immune system / SD1-3 / BA.2.12.1 / immune system / SD1-3 / BA.2.12.1 /  omicron variant / omicron variant /  virus / BA.2.86 virus / BA.2.86 | |||||||||||||||
| Function / homology |  Function and homology information Function and homology information virion component / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion ... virion component / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion ... virion component / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / virion component / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated endocytosis of virus by host cell / Attachment and Entry /  membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell /  receptor ligand activity / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / receptor ligand activity / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane /  viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||||||||
| Biological species |   Severe acute respiratory syndrome coronavirus 2 / Severe acute respiratory syndrome coronavirus 2 /   Homo sapiens (human) / Homo sapiens (human) /   Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) | |||||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.37 Å cryo EM / Resolution: 2.37 Å | |||||||||||||||
 Authors Authors | Duyvesteyn HME / Ren J / Stuart DI | |||||||||||||||
| Funding support |  United Kingdom, United Kingdom,  China, 4 items China, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: The SARS-CoV-2 neutralizing antibody response to SD1 and its evasion by BA.2.86. Authors: Daming Zhou / Piyada Supasa / Chang Liu / Aiste Dijokaite-Guraliuc / Helen M E Duyvesteyn / Muneeswaran Selvaraj / Alexander J Mentzer / Raksha Das / Wanwisa Dejnirattisai / Nigel Temperton ...Authors: Daming Zhou / Piyada Supasa / Chang Liu / Aiste Dijokaite-Guraliuc / Helen M E Duyvesteyn / Muneeswaran Selvaraj / Alexander J Mentzer / Raksha Das / Wanwisa Dejnirattisai / Nigel Temperton / Paul Klenerman / Susanna J Dunachie / Elizabeth E Fry / Juthathip Mongkolsapaya / Jingshan Ren / David I Stuart / Gavin R Screaton /    Abstract: Under pressure from neutralising antibodies induced by vaccination or infection the SARS-CoV-2 spike gene has become a hotspot for evolutionary change, leading to the failure of all mAbs developed ...Under pressure from neutralising antibodies induced by vaccination or infection the SARS-CoV-2 spike gene has become a hotspot for evolutionary change, leading to the failure of all mAbs developed for clinical use. Most potent antibodies bind to the receptor binding domain which has become heavily mutated. Here we study responses to a conserved epitope in sub-domain-1 (SD1) of spike which have become more prominent because of mutational escape from antibodies directed to the receptor binding domain. Some SD1 reactive mAbs show potent and broad neutralization of SARS-CoV-2 variants. We structurally map the dominant SD1 epitope and provide a mechanism of action by blocking interaction with ACE2. Mutations in SD1 have not been sustained to date, but one, E554K, leads to escape from mAbs. This mutation has now emerged in several sublineages including BA.2.86, reflecting selection pressure on the virus exerted by the increasing prominence of the anti-SD1 response. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18808.map.gz emd_18808.map.gz | 45.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18808-v30.xml emd-18808-v30.xml emd-18808.xml emd-18808.xml | 23.7 KB 23.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_18808.png emd_18808.png | 98.8 KB | ||
| Filedesc metadata |  emd-18808.cif.gz emd-18808.cif.gz | 7.9 KB | ||
| Others |  emd_18808_half_map_1.map.gz emd_18808_half_map_1.map.gz emd_18808_half_map_2.map.gz emd_18808_half_map_2.map.gz | 45 MB 45 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18808 http://ftp.pdbj.org/pub/emdb/structures/EMD-18808 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18808 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18808 | HTTPS FTP |
-Related structure data
| Related structure data |  8r1dMC  8cinC  8r1cC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18808.map.gz / Format: CCP4 / Size: 48.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18808.map.gz / Format: CCP4 / Size: 48.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM sharpened C3 symmetric map of SD1-6 fab in complex with BA.2.12.1 S. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.7303 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: CryoEM half map A of SD1-6 fab in complex with BA.2.12.1 S.
| File | emd_18808_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM half map A of SD1-6 fab in complex with BA.2.12.1 S. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: CryoEM half map B of SD1-6 fab in complex with BA.2.12.1 S.
| File | emd_18808_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM half map B of SD1-6 fab in complex with BA.2.12.1 S. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SD1-3 fab in complex with SARS-COV-2 BA.12.1 Spike Glycoprotein
| Entire | Name: SD1-3 fab in complex with SARS-COV-2 BA.12.1 Spike Glycoprotein |
|---|---|
| Components |
|
-Supramolecule #1: SD1-3 fab in complex with SARS-COV-2 BA.12.1 Spike Glycoprotein
| Supramolecule | Name: SD1-3 fab in complex with SARS-COV-2 BA.12.1 Spike Glycoprotein type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2 |
-Supramolecule #2: SD1-3 fab
| Supramolecule | Name: SD1-3 fab / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 Details: Sequence from antibody isolated from convalescent sera. |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: BA.2.12.1 Spike Glycoprotein Ectodomain
| Supramolecule | Name: BA.2.12.1 Spike Glycoprotein Ectodomain / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3 / Details: Has fibritin oligomerization tag |
|---|---|
| Source (natural) | Organism:   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2 |
-Macromolecule #1: SD1-3 Fab Heavy Chain
| Macromolecule | Name: SD1-3 Fab Heavy Chain / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.914263 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVESGGG LVQPGGSLRL SCTASGVTFS SYAMSWVRQA PGKGLEWVSG ISGSGDSTYY ADSVKGRFTI SRDNSKNTLY LQMNSLRVE DTAVYYCATH YCSGGSCPFD YWGQGTLVTV SS |
-Macromolecule #2: SD1-3 Fab Light Chain
| Macromolecule | Name: SD1-3 Fab Light Chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.218331 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: PSALTQPPSV SVSPGQTASI TCSGDKLGNK YAYWYQQKPG QSPVLVIFQD TMRPSGIPER FSASNSGNTA TLTISGTQSI DEADYYCQA WDSNTAVFGG GTKLTVL |
-Macromolecule #3: Spike glycoprotein,Fibritin
| Macromolecule | Name: Spike glycoprotein,Fibritin / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T4 (virus) Enterobacteria phage T4 (virus) |
| Molecular weight | Theoretical: 142.72925 KDa |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPV LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYYHKNNKSW MESEFRVYSS A NNCTFEYV ...String: MFVFLVLLPL VSSQCVNLIT RTQSYTNSFT RGVYYPDKVF RSSVLHSTQD LFLPFFSNVT WFHAIHVSGT NGTKRFDNPV LPFNDGVYF ASTEKSNIIR GWIFGTTLDS KTQSLLIVNN ATNVVIKVCE FQFCNDPFLD VYYHKNNKSW MESEFRVYSS A NNCTFEYV SQPFLMDLEG KQGNFKNLRE FVFKNIDGYF KIYSKHTPIN LGRDLPQGFS ALEPLVDLPI GINITRFQTL LA LHRSYLT PGDSSSGWTA GAAAYYVGYL QPRTFLLKYN ENGTITDAVD CALDPLSETK CTLKSFTVEK GIYQTSNFRV QPT ESIVRF PNITNLCPFD EVFNATRFAS VYAWNRKRIS NCVADYSVLY NFAPFFAFKC YGVSPTKLND LCFTNVYADS FVIR GNEVS QIAPGQTGNI ADYNYKLPDD FTGCVIAWNS NKLDSKVGGN YNYQYRLFRK SNLKPFERDI STEIYQAGNK PCNGV AGFN CYFPLRSYGF RPTYGVGHQP YRVVVLSFEL LHAPATVCGP KKSTNLVKNK CVNFNFNGLT GTGVLTESNK KFLPFQ QFG RDIADTTDAV RDPQTLEILD ITPCSFGGVS VITPGTNTSN QVAVLYQGVN CTEVPVAIHA DQLTPTWRVY STGSNVF QT RAGCLIGAEY VNNSYECDIP IGAGICASYQ TQTKSHRRAR SVASQSIIAY TMSLGAENLV AYSNNSIAIP TNFTISVT T EILPVSMTKT SVDCTMYICG DSTECSNLLL QYGSFCTQLK RALTGIAVEQ DKNTQEVFAQ VKQIYKTPPI KYFGGFNFS QILPDPSKPS KRSFIEDLLF NKVTLADAGF IKQYGDCLGD IAARDLICAQ KFNGLTVLPP LLTDEMIAQY TSALLAGTIT SGWTFGAGA ALQIPFAMQM AYRFNGIGVT QNVLYENQKL IANQFNSAIG KIQDSLSSTA SALGKLQDVV NHNAQALNTL V KQLSSKFG AISSVLNDIL SRLDKVEAEV QIDRLITGRL QSLQTYVTQQ LIRAAEIRAS ANLAATKMSE CVLGQSKRVD FC GKGYHLM SFPQSAPHGV VFLHVTYVPA QEKNFTTAPA ICHDGKAHFP REGVFVSNGT HWFVTQRNFY EPQIITTDNT FVS GNCDVV IGIVNNTVYD PLQPELDSFK EELDKYFKNH TSPDVDLGDI SGINASVVNI QKEIDRLNEV AKNLNESLID LQEL GKYEQ GSGYIPEAPR DGQAYVRKDG EWVLLSTFLG RSLEVLFQGP GHHHHHHHHG SAWSHPQFEK GGGSGGGSGG SAWSH PQFE K UniProtKB:  Spike glycoprotein, Fibritin Spike glycoprotein, Fibritin |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 30 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 43 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: C-flat-2/1 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.5 K / Instrument: FEI VITROBOT MARK IV |
| Details | Spike ectodomain incubated with SD1-3 fab in 2-fold molecular excess of sites (assuming three per trimeric spike unit). |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 165000 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 3282771 / Details: Blob picker in cryoSPARC live interface. |
|---|---|
| Startup model | Type of model: INSILICO MODEL In silico model: Ab initio model generated in cryoSPARC from 100,000 randomly selected picked particles. |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC |
| Final 3D classification | Number classes: 10 / Software - Name: cryoSPARC |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC |
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: C3 (3 fold cyclic ) / Resolution.type: BY AUTHOR / Resolution: 2.37 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 167816 ) / Resolution.type: BY AUTHOR / Resolution: 2.37 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 167816 |
 Movie
Movie Controller
Controller








 Z
Z Y
Y X
X