+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of mouse heavy-chain apoferritin | |||||||||
 Map data Map data | Main sharpened and masked map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  apoferritin / METAL BINDING PROTEIN apoferritin / METAL BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationIron uptake and transport / Golgi Associated Vesicle Biogenesis / iron ion sequestering activity /  autolysosome / autolysosome /  ferroxidase / intracellular sequestering of iron ion / ferroxidase / intracellular sequestering of iron ion /  ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen / Neutrophil degranulation ...Iron uptake and transport / Golgi Associated Vesicle Biogenesis / iron ion sequestering activity / ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen / Neutrophil degranulation ...Iron uptake and transport / Golgi Associated Vesicle Biogenesis / iron ion sequestering activity /  autolysosome / autolysosome /  ferroxidase / intracellular sequestering of iron ion / ferroxidase / intracellular sequestering of iron ion /  ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen / Neutrophil degranulation / ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen / Neutrophil degranulation /  ferric iron binding / ferric iron binding /  ferrous iron binding / iron ion transport / ferrous iron binding / iron ion transport /  immune response / iron ion binding / negative regulation of cell population proliferation / immune response / iron ion binding / negative regulation of cell population proliferation /  mitochondrion / extracellular region / identical protein binding / mitochondrion / extracellular region / identical protein binding /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Mus musculus (house mouse) Mus musculus (house mouse) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 1.5 Å cryo EM / Resolution: 1.5 Å | |||||||||
 Authors Authors | Dickerson JL / Russo CJ | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Ultramicroscopy / Year: 2024 Journal: Ultramicroscopy / Year: 2024Title: Accurate magnification determination for cryoEM using gold. Authors: Joshua L Dickerson / Erin Leahy / Mathew J Peet / Katerina Naydenova / Christopher J Russo /  Abstract: Determining the correct magnified pixel size of single-particle cryoEM micrographs is necessary to maximize resolution and enable accurate model building. Here we describe a simple and rapid ...Determining the correct magnified pixel size of single-particle cryoEM micrographs is necessary to maximize resolution and enable accurate model building. Here we describe a simple and rapid procedure for determining the absolute magnification in an electron cryomicroscope to a precision of <0.5%. We show how to use the atomic lattice spacings of crystals of thin and readily available test specimens, such as gold, as an absolute reference to determine magnification for both room temperature and cryogenic imaging. We compare this method to other commonly used methods, and show that it provides comparable accuracy in spite of its simplicity. This magnification calibration method provides a definitive reference quantity for data analysis and processing, simplifies the combination of multiple datasets from different microscopes and detectors, and improves the accuracy with which the contrast transfer function of the microscope can be determined. We also provide an open source program, magCalEM, which can be used to accurately estimate the magnified pixel size of a cryoEM dataset ex post facto. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17995.map.gz emd_17995.map.gz | 32.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17995-v30.xml emd-17995-v30.xml emd-17995.xml emd-17995.xml | 14.8 KB 14.8 KB | Display Display |  EMDB header EMDB header |
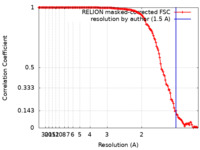
| FSC (resolution estimation) |  emd_17995_fsc.xml emd_17995_fsc.xml | 14 KB | Display |  FSC data file FSC data file |
| Images |  emd_17995.png emd_17995.png | 311 KB | ||
| Masks |  emd_17995_msk_1.map emd_17995_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17995.cif.gz emd-17995.cif.gz | 4.8 KB | ||
| Others |  emd_17995_half_map_1.map.gz emd_17995_half_map_1.map.gz emd_17995_half_map_2.map.gz emd_17995_half_map_2.map.gz | 189 MB 189 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17995 http://ftp.pdbj.org/pub/emdb/structures/EMD-17995 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17995 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17995 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17995.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17995.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main sharpened and masked map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.646 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17995_msk_1.map emd_17995_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_17995_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_17995_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mouse heavy-chain apoferritin
| Entire | Name: Mouse heavy-chain apoferritin |
|---|---|
| Components |
|
-Supramolecule #1: Mouse heavy-chain apoferritin
| Supramolecule | Name: Mouse heavy-chain apoferritin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Molecular weight | Theoretical: 485 KDa |
-Macromolecule #1: Apoferritin
| Macromolecule | Name: Apoferritin / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Mus musculus (house mouse) Mus musculus (house mouse) |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: PSQVRQNYHQ DAEAAINRQI NLELYASYVY LSMSCYFDRD DVALKNFAKY FLHQSHEERE HAEKLMKLQN QRGGRIFLQD IKKPDRDDW ESGLNAMECA LHLEKSVNQS LLELHKLATD KNDPHLCDFI ETYYLSEQVK SIKELGDHVT NLRKMGAPEA G MAEYLFDK HTLG UniProtKB:  Ferritin heavy chain Ferritin heavy chain |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10.8 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Material: GOLD / Support film - Material: GOLD / Support film - topology: HOLEY ARRAY / Support film - Film thickness: 50 / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.4000000000000001 µm / Nominal defocus min: 0.6 µm Bright-field microscopy / Nominal defocus max: 1.4000000000000001 µm / Nominal defocus min: 0.6 µm |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z
Z Y
Y X
X