+ Open data
Open data
- Basic information
Basic information
| Entry | 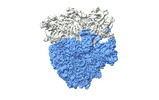 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
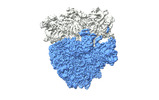
| Title | Cami1-ribosome local refinement map with mask on 50S subunit | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  CRISPR / cyclic oligoadenylate / CRISPR / cyclic oligoadenylate /  RNAse / RelE-toxin / RNAse / RelE-toxin /  RIBOSOME RIBOSOME | |||||||||
| Biological species |   Escherichia coli (E. coli) Escherichia coli (E. coli) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.9 Å cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Tamulaitiene G / Mogila I / Sasnauskas G / Tamulaitis G | |||||||||
| Funding support | Lithuania, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Ribosomal stalk-captured CARF-RelE ribonuclease inhibits translation following CRISPR signaling. Authors: Irmantas Mogila / Giedre Tamulaitiene / Konstanty Keda / Albertas Timinskas / Audrone Ruksenaite / Giedrius Sasnauskas / Česlovas Venclovas / Virginijus Siksnys / Gintautas Tamulaitis Abstract: Prokaryotic type III CRISPR-Cas antiviral systems employ cyclic oligoadenylate (cA) signaling to activate a diverse range of auxiliary proteins that reinforce the CRISPR-Cas defense. Here we ...Prokaryotic type III CRISPR-Cas antiviral systems employ cyclic oligoadenylate (cA) signaling to activate a diverse range of auxiliary proteins that reinforce the CRISPR-Cas defense. Here we characterize a class of cA-dependent effector proteins named CRISPR-Cas-associated messenger RNA (mRNA) interferase 1 (Cami1) consisting of a CRISPR-associated Rossmann fold sensor domain fused to winged helix-turn-helix and a RelE-family mRNA interferase domain. Upon activation by cyclic tetra-adenylate (cA), Cami1 cleaves mRNA exposed at the ribosomal A-site thereby depleting mRNA and leading to cell growth arrest. The structures of apo-Cami1 and the ribosome-bound Cami1-cA complex delineate the conformational changes that lead to Cami1 activation and the mechanism of Cami1 binding to a bacterial ribosome, revealing unexpected parallels with eukaryotic ribosome-inactivating proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17666.map.gz emd_17666.map.gz | 123.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17666-v30.xml emd-17666-v30.xml emd-17666.xml emd-17666.xml | 13.3 KB 13.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17666_fsc.xml emd_17666_fsc.xml | 18.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_17666.png emd_17666.png | 75.6 KB | ||
| Masks |  emd_17666_msk_1.map emd_17666_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17666.cif.gz emd-17666.cif.gz | 4 KB | ||
| Others |  emd_17666_half_map_1.map.gz emd_17666_half_map_1.map.gz emd_17666_half_map_2.map.gz emd_17666_half_map_2.map.gz | 227 MB 227 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17666 http://ftp.pdbj.org/pub/emdb/structures/EMD-17666 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17666 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17666 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_17666.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17666.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17666_msk_1.map emd_17666_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
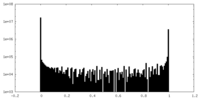
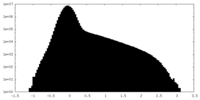
| Density Histograms |
-Half map: #2
| File | emd_17666_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_17666_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cami1 bound in 70S E.coli ribosome
| Entire | Name: Cami1 bound in 70S E.coli ribosome |
|---|---|
| Components |
|
-Supramolecule #1: Cami1 bound in 70S E.coli ribosome
| Supramolecule | Name: Cami1 bound in 70S E.coli ribosome / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Average electron dose: 30.0 e/Å2 |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X