+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1566 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
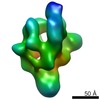
| Title | Activator bound RNA polymerase holoenzyme | |||||||||
 Map data Map data | Activator bound RNA polymerase holoenzyme | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA polymerase / RNAP / Sigma 54 / PspF / transcription initiation / AAA ATPase / sigma factor | |||||||||
| Biological species |   Klebsiella pneumoniae (bacteria) Klebsiella pneumoniae (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 21.0 Å | |||||||||
 Authors Authors | Bose D / Pape T / Burrows PC / Rappas M / Wigneshweraraj SR / Buck M / Zhang X | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2008 Journal: Mol Cell / Year: 2008Title: Organization of an activator-bound RNA polymerase holoenzyme. Authors: Daniel Bose / Tillmann Pape / Patricia C Burrows / Mathieu Rappas / Siva R Wigneshweraraj / Martin Buck / Xiaodong Zhang /  Abstract: Transcription initiation involves the conversion from closed promoter complexes, comprising RNA polymerase (RNAP) and double-stranded promoter DNA, to open complexes, in which the enzyme is able to ...Transcription initiation involves the conversion from closed promoter complexes, comprising RNA polymerase (RNAP) and double-stranded promoter DNA, to open complexes, in which the enzyme is able to access the DNA template in a single-stranded form. The complex between bacterial RNAP and its major variant sigma factor sigma(54) remains as a closed complex until ATP hydrolysis-dependent remodeling by activator proteins occurs. This remodeling facilitates DNA melting and allows the transition to the open complex. Here we present cryoelectron microscopy reconstructions of bacterial RNAP in complex with sigma(54) alone, and of RNAP-sigma(54) with an AAA+ activator. Together with photo-crosslinking data that establish the location of promoter DNA within the complexes, we explain why the RNAP-sigma(54) closed complex is unable to access the DNA template and propose how the structural changes induced by activator binding can initiate conformational changes that ultimately result in formation of the open complex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1566.map.gz emd_1566.map.gz | 7.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1566-v30.xml emd-1566-v30.xml emd-1566.xml emd-1566.xml | 12 KB 12 KB | Display Display |  EMDB header EMDB header |
| Images |  1566.gif 1566.gif | 31.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1566 http://ftp.pdbj.org/pub/emdb/structures/EMD-1566 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1566 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1566 | HTTPS FTP |
-Validation report
| Summary document |  emd_1566_validation.pdf.gz emd_1566_validation.pdf.gz | 194.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1566_full_validation.pdf.gz emd_1566_full_validation.pdf.gz | 193.4 KB | Display | |
| Data in XML |  emd_1566_validation.xml.gz emd_1566_validation.xml.gz | 5.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1566 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1566 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1566 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1566 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1566.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1566.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Activator bound RNA polymerase holoenzyme | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Complex of E.coli RNA polymerase, Sigma 54 and PspF1-275 trapped ...
| Entire | Name: Complex of E.coli RNA polymerase, Sigma 54 and PspF1-275 trapped with ADP.AlFx |
|---|---|
| Components |
|
-Supramolecule #1000: Complex of E.coli RNA polymerase, Sigma 54 and PspF1-275 trapped ...
| Supramolecule | Name: Complex of E.coli RNA polymerase, Sigma 54 and PspF1-275 trapped with ADP.AlFx type: sample / ID: 1000 / Details: All components are present on SDS-PAGE. Oligomeric state: monomer of RNAP sigma 54 holoenzyme bound to hexamer of PspF Number unique components: 3 |
|---|---|
| Molecular weight | Theoretical: 630 KDa |
-Macromolecule #1: RNA polymerase
| Macromolecule | Name: RNA polymerase / type: protein_or_peptide / ID: 1 / Name.synonym: RNAP / Number of copies: 1 / Oligomeric state: pentamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 390 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: Sigma 54
| Macromolecule | Name: Sigma 54 / type: protein_or_peptide / ID: 2 / Name.synonym: s45 / Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Klebsiella pneumoniae (bacteria) Klebsiella pneumoniae (bacteria) |
| Molecular weight | Theoretical: 54 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #3: Phage shock protein F
| Macromolecule | Name: Phage shock protein F / type: protein_or_peptide / ID: 3 / Name.synonym: PspF / Number of copies: 1 / Oligomeric state: hexamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 180 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 10mM Tris-HCl pH8.0, 150mM NaCl, 10mM MgCl2, |
|---|---|
| Grid | Details: Copper 400 mesh |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 83 K / Instrument: OTHER / Details: Vitrification instrument: Vitrobot |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG/UT |
|---|---|
| Temperature | Average: 91 K |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 6.35 µm / Number real images: 50 / Average electron dose: 10 e/Å2 / Od range: 4.8 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 48700 / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.1 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: particles |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 21.0 Å / Resolution method: OTHER / Software - Name: Imagic V / Number images used: 19581 |
| Final two d classification | Number classes: 695 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: Situs |
| Details | Protocol: Rigid body. Structure positioned by hand and fitting refined in Situs |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 0.6 |
-Atomic model buiding 2
| Initial model | PDB ID: |
|---|---|
| Software | Name: Situs |
| Details | Protocol: Rigid body. Structure positioned by hand and fitting refined in Situs |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller