+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human TSEN with pre-tRNA-Tyr-GTA | |||||||||
 Map data Map data | map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNP /  endonuclease / endonuclease /  tRNA / tRNA /  splicing / splicing /  RNA BINDING PROTEIN RNA BINDING PROTEIN | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.8 Å cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Sekulovski S / Trowitzsch S | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structural basis of substrate recognition by human tRNA splicing endonuclease TSEN. Authors: Samoil Sekulovski / Lukas Sušac / Lukas S Stelzl / Robert Tampé / Simon Trowitzsch /  Abstract: Heterotetrameric human transfer RNA (tRNA) splicing endonuclease TSEN catalyzes intron excision from precursor tRNAs (pre-tRNAs), utilizing two composite active sites. Mutations in TSEN and its ...Heterotetrameric human transfer RNA (tRNA) splicing endonuclease TSEN catalyzes intron excision from precursor tRNAs (pre-tRNAs), utilizing two composite active sites. Mutations in TSEN and its associated RNA kinase CLP1 are linked to the neurodegenerative disease pontocerebellar hypoplasia (PCH). Despite the essential function of TSEN, the three-dimensional assembly of TSEN-CLP1, the mechanism of substrate recognition, and the structural consequences of disease mutations are not understood in molecular detail. Here, we present single-particle cryogenic electron microscopy reconstructions of human TSEN with intron-containing pre-tRNAs. TSEN recognizes the body of pre-tRNAs and pre-positions the 3' splice site for cleavage by an intricate protein-RNA interaction network. TSEN subunits exhibit large unstructured regions flexibly tethering CLP1. Disease mutations localize far from the substrate-binding interface and destabilize TSEN. Our work delineates molecular principles of pre-tRNA recognition and cleavage by human TSEN and rationalizes mutations associated with PCH. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14925.map.gz emd_14925.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14925-v30.xml emd-14925-v30.xml emd-14925.xml emd-14925.xml | 11.8 KB 11.8 KB | Display Display |  EMDB header EMDB header |
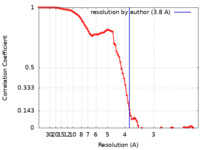
| FSC (resolution estimation) |  emd_14925_fsc.xml emd_14925_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_14925.png emd_14925.png | 46.7 KB | ||
| Masks |  emd_14925_msk_1.map emd_14925_msk_1.map | 64 MB |  Mask map Mask map | |
| Others |  emd_14925_half_map_1.map.gz emd_14925_half_map_1.map.gz emd_14925_half_map_2.map.gz emd_14925_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14925 http://ftp.pdbj.org/pub/emdb/structures/EMD-14925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14925 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14925 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14925.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14925.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0635 Å | ||||||||||||||||||||||||||||||||||||
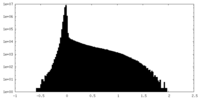
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14925_msk_1.map emd_14925_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
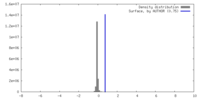
| Density Histograms |
-Half map: half map A
| File | emd_14925_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half_map_A | ||||||||||||
| Projections & Slices |
| ||||||||||||
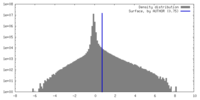
| Density Histograms |
-Half map: half map B
| File | emd_14925_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half_map_B | ||||||||||||
| Projections & Slices |
| ||||||||||||
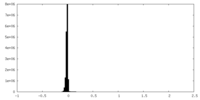
| Density Histograms |
- Sample components
Sample components
-Entire : Human TSEN with pre-tRNA-Tyr-GTA
| Entire | Name: Human TSEN with pre-tRNA-Tyr-GTA |
|---|---|
| Components |
|
-Supramolecule #1: Human TSEN with pre-tRNA-Tyr-GTA
| Supramolecule | Name: Human TSEN with pre-tRNA-Tyr-GTA / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 81.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)