+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13855 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of respiratory syncytial virus matrix layer | ||||||||||||||||||
 Map data Map data | Sub-tomogram average of the envelope and matrix layer of respiratory syncytial virus | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Matrix /  VIRAL PROTEIN VIRAL PROTEIN | ||||||||||||||||||
| Biological species |   Respiratory syncytial virus A2 Respiratory syncytial virus A2 | ||||||||||||||||||
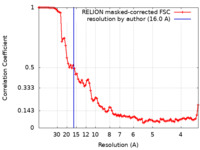
| Method | subtomogram averaging /  cryo EM / Resolution: 16.0 Å cryo EM / Resolution: 16.0 Å | ||||||||||||||||||
 Authors Authors | Conley MJ / Vijayakrishnan S / Bhella D | ||||||||||||||||||
| Funding support |  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: EMBO J / Year: 2022 Journal: EMBO J / Year: 2022Title: Helical ordering of envelope-associated proteins and glycoproteins in respiratory syncytial virus. Authors: Michaela J Conley / Judith M Short / Andrew M Burns / James Streetley / Joshua Hutchings / Saskia E Bakker / B Joanne Power / Hussain Jaffery / Joanne Haney / Giulia Zanetti / Pablo R Murcia ...Authors: Michaela J Conley / Judith M Short / Andrew M Burns / James Streetley / Joshua Hutchings / Saskia E Bakker / B Joanne Power / Hussain Jaffery / Joanne Haney / Giulia Zanetti / Pablo R Murcia / Murray Stewart / Rachel Fearns / Swetha Vijayakrishnan / David Bhella /   Abstract: Human respiratory syncytial virus (RSV) causes severe respiratory illness in children and the elderly. Here, using cryogenic electron microscopy and tomography combined with computational image ...Human respiratory syncytial virus (RSV) causes severe respiratory illness in children and the elderly. Here, using cryogenic electron microscopy and tomography combined with computational image analysis and three-dimensional reconstruction, we show that there is extensive helical ordering of the envelope-associated proteins and glycoproteins of RSV filamentous virions. We calculated a 16 Å resolution sub-tomogram average of the matrix protein (M) layer that forms an endoskeleton below the viral envelope. These data define a helical lattice of M-dimers, showing how M is oriented relative to the viral envelope. Glycoproteins that stud the viral envelope were also found to be helically ordered, a property that was coordinated by the M-layer. Furthermore, envelope glycoproteins clustered in pairs, a feature that may have implications for the conformation of fusion (F) glycoprotein epitopes that are the principal target for vaccine and monoclonal antibody development. We also report the presence, in authentic virus infections, of N-RNA rings packaged within RSV virions. These data provide molecular insight into the organisation of the virion and the mechanism of its assembly. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13855.map.gz emd_13855.map.gz | 56.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13855-v30.xml emd-13855-v30.xml emd-13855.xml emd-13855.xml | 17.3 KB 17.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13855_fsc.xml emd_13855_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_13855.png emd_13855.png | 114.6 KB | ||
| Masks |  emd_13855_msk_1.map emd_13855_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13855.cif.gz emd-13855.cif.gz | 5.3 KB | ||
| Others |  emd_13855_half_map_1.map.gz emd_13855_half_map_1.map.gz emd_13855_half_map_2.map.gz emd_13855_half_map_2.map.gz | 57.6 MB 57.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13855 http://ftp.pdbj.org/pub/emdb/structures/EMD-13855 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13855 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13855 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13855.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13855.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sub-tomogram average of the envelope and matrix layer of respiratory syncytial virus | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.786 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
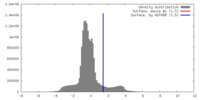
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_13855_msk_1.map emd_13855_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
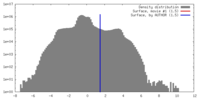
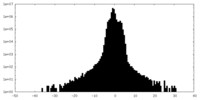
| Density Histograms |
-Half map: Half-map 2 (not gold-standard)
| File | emd_13855_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 (not gold-standard) | ||||||||||||
| Projections & Slices |
| ||||||||||||
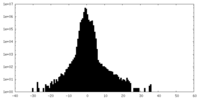
| Density Histograms |
-Half map: Half-map 1 (not gold-standard)
| File | emd_13855_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 (not gold-standard) | ||||||||||||
| Projections & Slices |
| ||||||||||||
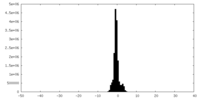
| Density Histograms |
- Sample components
Sample components
-Entire : Matrix layer of respiratory syncytial virus
| Entire | Name: Matrix layer of respiratory syncytial virus |
|---|---|
| Components |
|
-Supramolecule #1: Matrix layer of respiratory syncytial virus
| Supramolecule | Name: Matrix layer of respiratory syncytial virus / type: complex / ID: 1 / Parent: 0 Details: Sub tomogram averaging was performed focusing on the envelope of filamentous virions propagated directly on the cryo-EM grid. |
|---|---|
| Source (natural) | Organism:   Respiratory syncytial virus A2 Respiratory syncytial virus A2 |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
| Details | Virions were propagated in A549 cells, grown on gold quantifoil cryo-EM grids that had been pre-treated with laminin. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Average exposure time: 1.0 sec. / Average electron dose: 2.0 e/Å2 Details: Data collected at the UK electron bioimaging centre at Diamond Light Source (eBIC) on a Thermo-Fisher Scientific Titan Krios microscope equipped with a Gatan BioQuantum K2 energy filtered ...Details: Data collected at the UK electron bioimaging centre at Diamond Light Source (eBIC) on a Thermo-Fisher Scientific Titan Krios microscope equipped with a Gatan BioQuantum K2 energy filtered direct detection camera. Dose-symmetric tilt-series collection was performed using SerialEM. Energy-filtered images were collected with a slit-width of 20eV and an applied defocus of between -2 and -4.5 microns. One second exposures were recorded with an electron exposure of 2 electrons/A2, partitioned over five movie frames. A total of 41 images were recorded per tilt-series at 3 degree intervals from -60 to +60, thus a total exposure of 82 electrons/A2 was applied per tilt-series. Tilt-series were recorded at a nominal column magnification of 81k corresponding to a calibrated pixel size of 1.79 A at the specimen scale. |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)