[English] 日本語
 Yorodumi
Yorodumi- EMDB-13577: Alpha-synuclein amyloid fibres incubated with DNAJB1, Hsc70, and ATP -
+ Open data
Open data
- Basic information
Basic information
| Entry | 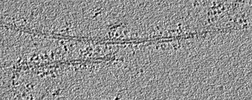 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Alpha-synuclein amyloid fibres incubated with DNAJB1, Hsc70, and ATP | |||||||||
 Map data Map data | Alpha-synuclein incubated with DNAJB1, Hsc70, and ATP. Small clusters of chaperones bound to the fibres are interspersed with sparse decoration. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Monistrol J / Saibil HR | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2022 Journal: EMBO J / Year: 2022Title: Cooperative amyloid fibre binding and disassembly by the Hsp70 disaggregase. Authors: Joseph George Beton / Jim Monistrol / Anne Wentink / Erin C Johnston / Anthony John Roberts / Bernd Gerhard Bukau / Bart W Hoogenboom / Helen R Saibil /   Abstract: Although amyloid fibres are highly stable protein aggregates, a specific combination of human Hsp70 system chaperones can disassemble them, including fibres formed of α-synuclein, huntingtin, or Tau. ...Although amyloid fibres are highly stable protein aggregates, a specific combination of human Hsp70 system chaperones can disassemble them, including fibres formed of α-synuclein, huntingtin, or Tau. Disaggregation requires the ATPase activity of the constitutively expressed Hsp70 family member, Hsc70, together with the J domain protein DNAJB1 and the nucleotide exchange factor Apg2. Clustering of Hsc70 on the fibrils appears to be necessary for disassembly. Here we use atomic force microscopy to show that segments of in vitro assembled α-synuclein fibrils are first coated with chaperones and then undergo bursts of rapid, unidirectional disassembly. Cryo-electron tomography and total internal reflection fluorescence microscopy reveal fibrils with regions of densely bound chaperones, preferentially at one end of the fibre. Sub-stoichiometric amounts of Apg2 relative to Hsc70 dramatically increase recruitment of Hsc70 to the fibres, creating localised active zones that then undergo rapid disassembly at a rate of ~ 4 subunits per second. The observed unidirectional bursts of Hsc70 loading and unravelling may be explained by differences between the two ends of the polar fibre structure. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13577.map.gz emd_13577.map.gz | 1.1 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13577-v30.xml emd-13577-v30.xml emd-13577.xml emd-13577.xml | 11.4 KB 11.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13577.png emd_13577.png | 102.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13577 http://ftp.pdbj.org/pub/emdb/structures/EMD-13577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13577 | HTTPS FTP |
-Validation report
| Summary document |  emd_13577_validation.pdf.gz emd_13577_validation.pdf.gz | 366.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_13577_full_validation.pdf.gz emd_13577_full_validation.pdf.gz | 365.7 KB | Display | |
| Data in XML |  emd_13577_validation.xml.gz emd_13577_validation.xml.gz | 5.1 KB | Display | |
| Data in CIF |  emd_13577_validation.cif.gz emd_13577_validation.cif.gz | 5.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13577 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13577 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13577 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13577 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_13577.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13577.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Alpha-synuclein incubated with DNAJB1, Hsc70, and ATP. Small clusters of chaperones bound to the fibres are interspersed with sparse decoration. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 8.52 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Complex of alpha-synuclein amyloid fibres, DNAJB1 and Hsc70 chaperones
| Entire | Name: Complex of alpha-synuclein amyloid fibres, DNAJB1 and Hsc70 chaperones |
|---|---|
| Components |
|
-Supramolecule #1: Complex of alpha-synuclein amyloid fibres, DNAJB1 and Hsc70 chaperones
| Supramolecule | Name: Complex of alpha-synuclein amyloid fibres, DNAJB1 and Hsc70 chaperones type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: C-flat-1.2/1.3 / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY | ||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE | ||||||||||||||||||||||||
| Sectioning | Other: NO SECTIONING | ||||||||||||||||||||||||
| Fiducial marker | Manufacturer: EMS / Diameter: 10 nm |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 150.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Software - Name:  IMOD (ver. 4.9.0) / Number images used: 41 IMOD (ver. 4.9.0) / Number images used: 41 |
|---|
 Movie
Movie Controller
Controller




