[English] 日本語
 Yorodumi
Yorodumi- EMDB-1322: Insights into transcription initiation and termination from the e... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1322 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
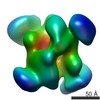
| Title | Insights into transcription initiation and termination from the electron microscopy structure of yeast RNA polymerase III. | |||||||||
 Map data Map data | Three volume calculated by back projection | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 17.0 Å | |||||||||
 Authors Authors | Fernandez-Tornero C / Bottcher B / Riva M / Carles C / Steuerwald U / Ruigrok RW / Sentenac A / Muller CW / Schoehn G | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2007 Journal: Mol Cell / Year: 2007Title: Insights into transcription initiation and termination from the electron microscopy structure of yeast RNA polymerase III. Authors: Carlos Fernández-Tornero / Bettina Böttcher / Michel Riva / Christophe Carles / Ulrich Steuerwald / Rob W H Ruigrok / André Sentenac / Christoph W Müller / Guy Schoehn /  Abstract: RNA polymerase III (RNAPIII) synthesizes tRNA, 5S RNA, U6 snRNA, and other small RNAs. The structure of yeast RNAPIII, determined at 17 A resolution by cryo-electron microscopy and single-particle ...RNA polymerase III (RNAPIII) synthesizes tRNA, 5S RNA, U6 snRNA, and other small RNAs. The structure of yeast RNAPIII, determined at 17 A resolution by cryo-electron microscopy and single-particle analysis, reveals a hand-like shape typical of RNA polymerases. Compared to RNAPII, RNAPIII is characterized by a bulkier stalk and by prominent features extending from the DNA binding cleft. We attribute the latter primarily to five RNAPIII-specific subunits, present as two distinct subcomplexes (C82/C34/C31 and C53/C37). Antibody labeling experiments localize the C82/C34/C31 subcomplex to the clamp side of the DNA binding cleft, consistent with its known role in transcription initiation. The C53/C37 subcomplex appears to be situated across the cleft, near the presumed location of downstream DNA, accounting for its role in transcription termination. Our structure rationalizes available mutagenesis and biochemical data and provides insights into RNAPIII-mediated transcription. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1322.map.gz emd_1322.map.gz | 2.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1322-v30.xml emd-1322-v30.xml emd-1322.xml emd-1322.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
| Images |  1322.gif 1322.gif | 15.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1322 http://ftp.pdbj.org/pub/emdb/structures/EMD-1322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1322 | HTTPS FTP |
-Validation report
| Summary document |  emd_1322_validation.pdf.gz emd_1322_validation.pdf.gz | 201.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1322_full_validation.pdf.gz emd_1322_full_validation.pdf.gz | 200.6 KB | Display | |
| Data in XML |  emd_1322_validation.xml.gz emd_1322_validation.xml.gz | 5.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1322 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1322 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1322 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1322 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1322.map.gz / Format: CCP4 / Size: 11.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1322.map.gz / Format: CCP4 / Size: 11.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Three volume calculated by back projection | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.75 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : yeast RNA polymerase III
+Supramolecule #1000: yeast RNA polymerase III
+Macromolecule #1: C160
+Macromolecule #2: C128
+Macromolecule #3: AC40
+Macromolecule #4: AC19
+Macromolecule #5: ABC27
+Macromolecule #6: ABC23
+Macromolecule #7: ABC14.5
+Macromolecule #8: ABC10alpha
+Macromolecule #9: ABC10beta
+Macromolecule #10: C25
+Macromolecule #11: C17
+Macromolecule #12: C53
+Macromolecule #13: C37
+Macromolecule #14: C11
+Macromolecule #15: C82
+Macromolecule #16: C34
+Macromolecule #17: C31
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM Tris pH 8.0, 200 mM Ammonium Sulfate, 10 mM DTT |
| Staining | Type: NEGATIVE / Details: coventional Cryo EM |
| Grid | Details: home made holey grids |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 100 K / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: modified controlled environment vitrification Method: Manual blot for 1 second before plunging |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010F |
|---|---|
| Temperature | Average: 100 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 100,000 |
| Date | Jan 1, 2005 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 12 / Average electron dose: 10 e/Å2 / Od range: 1 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 40000 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 1.4 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.8 µm / Nominal magnification: 40000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: OTHER |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 17.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: spider imagic Details: Projection matching started with a random conical tilt model Number images used: 10000 |
| Final angle assignment | Details: 386 reprojections |
| Final two d classification | Number classes: 386 |
-Atomic model buiding 1
| Software | Name: Situs |
|---|---|
| Details | Protocol: Rigid Body |
| Refinement | Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller