[English] 日本語
 Yorodumi
Yorodumi- EMDB-0510: Protein Phosphatase 2A (Aalpha-B56alpha-Calpha) holoenzyme in com... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0510 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Protein Phosphatase 2A (Aalpha-B56alpha-Calpha) holoenzyme in complex with a Small Molecule Activator of PP2A (SMAP) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  Holoenzyme complex / Holoenzyme complex /  Phosphatase / Activator / HYDROLASE-ACTIVATOR complex Phosphatase / Activator / HYDROLASE-ACTIVATOR complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of lipid kinase activity / regulation of protein autophosphorylation / meiotic spindle elongation / Integration of energy metabolism / PP2A-mediated dephosphorylation of key metabolic factors /  regulation of microtubule binding / MASTL Facilitates Mitotic Progression / mitotic sister chromatid separation / regulation of meiotic cell cycle process involved in oocyte maturation / protein phosphatase type 2A complex ...negative regulation of lipid kinase activity / regulation of protein autophosphorylation / meiotic spindle elongation / Integration of energy metabolism / PP2A-mediated dephosphorylation of key metabolic factors / regulation of microtubule binding / MASTL Facilitates Mitotic Progression / mitotic sister chromatid separation / regulation of meiotic cell cycle process involved in oocyte maturation / protein phosphatase type 2A complex ...negative regulation of lipid kinase activity / regulation of protein autophosphorylation / meiotic spindle elongation / Integration of energy metabolism / PP2A-mediated dephosphorylation of key metabolic factors /  regulation of microtubule binding / MASTL Facilitates Mitotic Progression / mitotic sister chromatid separation / regulation of meiotic cell cycle process involved in oocyte maturation / protein phosphatase type 2A complex / meiotic sister chromatid cohesion, centromeric / peptidyl-serine dephosphorylation / peptidyl-threonine dephosphorylation / regulation of microtubule binding / MASTL Facilitates Mitotic Progression / mitotic sister chromatid separation / regulation of meiotic cell cycle process involved in oocyte maturation / protein phosphatase type 2A complex / meiotic sister chromatid cohesion, centromeric / peptidyl-serine dephosphorylation / peptidyl-threonine dephosphorylation /  : / positive regulation of microtubule binding / negative regulation of tyrosine phosphorylation of STAT protein / Regulation of glycolysis by fructose 2,6-bisphosphate metabolism / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / female meiotic nuclear division / protein antigen binding / protein phosphatase regulator activity / ceramide metabolic process / : / positive regulation of microtubule binding / negative regulation of tyrosine phosphorylation of STAT protein / Regulation of glycolysis by fructose 2,6-bisphosphate metabolism / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / female meiotic nuclear division / protein antigen binding / protein phosphatase regulator activity / ceramide metabolic process /  GABA receptor binding / negative regulation of epithelial to mesenchymal transition / APC truncation mutants have impaired AXIN binding / AXIN missense mutants destabilize the destruction complex / Truncations of AMER1 destabilize the destruction complex / Initiation of Nuclear Envelope (NE) Reformation / ERKs are inactivated / positive regulation of extrinsic apoptotic signaling pathway in absence of ligand / Beta-catenin phosphorylation cascade / Signaling by GSK3beta mutants / CTNNB1 S33 mutants aren't phosphorylated / CTNNB1 S37 mutants aren't phosphorylated / CTNNB1 S45 mutants aren't phosphorylated / CTNNB1 T41 mutants aren't phosphorylated / GABA receptor binding / negative regulation of epithelial to mesenchymal transition / APC truncation mutants have impaired AXIN binding / AXIN missense mutants destabilize the destruction complex / Truncations of AMER1 destabilize the destruction complex / Initiation of Nuclear Envelope (NE) Reformation / ERKs are inactivated / positive regulation of extrinsic apoptotic signaling pathway in absence of ligand / Beta-catenin phosphorylation cascade / Signaling by GSK3beta mutants / CTNNB1 S33 mutants aren't phosphorylated / CTNNB1 S37 mutants aren't phosphorylated / CTNNB1 S45 mutants aren't phosphorylated / CTNNB1 T41 mutants aren't phosphorylated /  M band / M band /  regulation of Wnt signaling pathway / negative regulation of protein localization to plasma membrane / Disassembly of the destruction complex and recruitment of AXIN to the membrane / regulation of growth / negative regulation of glycolytic process through fructose-6-phosphate / positive regulation of NLRP3 inflammasome complex assembly / myosin phosphatase activity / regulation of Wnt signaling pathway / negative regulation of protein localization to plasma membrane / Disassembly of the destruction complex and recruitment of AXIN to the membrane / regulation of growth / negative regulation of glycolytic process through fructose-6-phosphate / positive regulation of NLRP3 inflammasome complex assembly / myosin phosphatase activity /  protein serine/threonine phosphatase activity / CTLA4 inhibitory signaling / Platelet sensitization by LDL / negative regulation of MAPK cascade / protein-serine/threonine phosphatase / protein serine/threonine phosphatase activity / CTLA4 inhibitory signaling / Platelet sensitization by LDL / negative regulation of MAPK cascade / protein-serine/threonine phosphatase /  regulation of cell differentiation / T cell homeostasis / ERK/MAPK targets / protein phosphatase activator activity / regulation of G1/S transition of mitotic cell cycle / regulation of cell differentiation / T cell homeostasis / ERK/MAPK targets / protein phosphatase activator activity / regulation of G1/S transition of mitotic cell cycle /  phosphoprotein phosphatase activity / phosphoprotein phosphatase activity /  regulation of DNA replication / mesoderm development / regulation of DNA replication / mesoderm development /  chromosome, centromeric region / DARPP-32 events / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / lateral plasma membrane / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / chromosome, centromeric region / DARPP-32 events / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / lateral plasma membrane / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal /  regulation of cell adhesion / Cyclin A/B1/B2 associated events during G2/M transition / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / positive regulation of protein dephosphorylation / Loss of Nlp from mitotic centrosomes / Loss of proteins required for interphase microtubule organization from the centrosome / Recruitment of mitotic centrosome proteins and complexes / Resolution of Sister Chromatid Cohesion / Recruitment of NuMA to mitotic centrosomes / Anchoring of the basal body to the plasma membrane / AURKA Activation by TPX2 / protein dephosphorylation / regulation of cell adhesion / Cyclin A/B1/B2 associated events during G2/M transition / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / positive regulation of protein dephosphorylation / Loss of Nlp from mitotic centrosomes / Loss of proteins required for interphase microtubule organization from the centrosome / Recruitment of mitotic centrosome proteins and complexes / Resolution of Sister Chromatid Cohesion / Recruitment of NuMA to mitotic centrosomes / Anchoring of the basal body to the plasma membrane / AURKA Activation by TPX2 / protein dephosphorylation /  RNA splicing / meiotic cell cycle / response to organic substance / RNA splicing / meiotic cell cycle / response to organic substance /  protein tyrosine phosphatase activity / protein tyrosine phosphatase activity /  chromosome segregation / RHO GTPases Activate Formins / response to lead ion / chromosome segregation / RHO GTPases Activate Formins / response to lead ion /  regulation of protein phosphorylation / Spry regulation of FGF signaling / RAF activation / PKR-mediated signaling / Degradation of beta-catenin by the destruction complex / tau protein binding / positive regulation of protein serine/threonine kinase activity / negative regulation of cell growth / Z disc / regulation of protein phosphorylation / Spry regulation of FGF signaling / RAF activation / PKR-mediated signaling / Degradation of beta-catenin by the destruction complex / tau protein binding / positive regulation of protein serine/threonine kinase activity / negative regulation of cell growth / Z disc /  kinase binding / kinase binding /  spindle pole / Negative regulation of MAPK pathway / Separation of Sister Chromatids / Cyclin D associated events in G1 / microtubule cytoskeleton / spindle pole / Negative regulation of MAPK pathway / Separation of Sister Chromatids / Cyclin D associated events in G1 / microtubule cytoskeleton /  Regulation of PLK1 Activity at G2/M Transition / Regulation of TP53 Degradation Regulation of PLK1 Activity at G2/M Transition / Regulation of TP53 DegradationSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
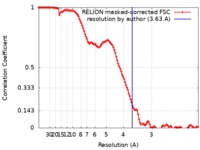
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.63 Å cryo EM / Resolution: 3.63 Å | |||||||||
 Authors Authors | Huang W / Taylor D | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: Selective PP2A Enhancement through Biased Heterotrimer Stabilization. Authors: Daniel Leonard / Wei Huang / Sudeh Izadmehr / Caitlin M O'Connor / Danica D Wiredja / Zhizhi Wang / Nilesh Zaware / Yinghua Chen / Daniela M Schlatzer / Janna Kiselar / Nikhil Vasireddi / ...Authors: Daniel Leonard / Wei Huang / Sudeh Izadmehr / Caitlin M O'Connor / Danica D Wiredja / Zhizhi Wang / Nilesh Zaware / Yinghua Chen / Daniela M Schlatzer / Janna Kiselar / Nikhil Vasireddi / Stefan Schüchner / Abbey L Perl / Matthew D Galsky / Wenqing Xu / David L Brautigan / Egon Ogris / Derek J Taylor / Goutham Narla /   Abstract: Impairment of protein phosphatases, including the family of serine/threonine phosphatases designated PP2A, is essential for the pathogenesis of many diseases, including cancer. The ability of PP2A to ...Impairment of protein phosphatases, including the family of serine/threonine phosphatases designated PP2A, is essential for the pathogenesis of many diseases, including cancer. The ability of PP2A to dephosphorylate hundreds of proteins is regulated by over 40 specificity-determining regulatory "B" subunits that compete for assembly and activation of heterogeneous PP2A heterotrimers. Here, we reveal how a small molecule, DT-061, specifically stabilizes the B56α-PP2A holoenzyme in a fully assembled, active state to dephosphorylate selective substrates, such as its well-known oncogenic target, c-Myc. Our 3.6 Å structure identifies molecular interactions between DT-061 and all three PP2A subunits that prevent dissociation of the active enzyme and highlight inherent mechanisms of PP2A complex assembly. Thus, our findings provide fundamental insights into PP2A complex assembly and regulation, identify a unique interfacial stabilizing mode of action for therapeutic targeting, and aid in the development of phosphatase-based therapeutics tailored against disease specific phospho-protein targets. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0510.map.gz emd_0510.map.gz | 69.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0510-v30.xml emd-0510-v30.xml emd-0510.xml emd-0510.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0510_fsc.xml emd_0510_fsc.xml | 12 KB | Display |  FSC data file FSC data file |
| Images |  emd_0510.png emd_0510.png | 255.7 KB | ||
| Masks |  emd_0510_msk_1.map emd_0510_msk_1.map | 144.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-0510.cif.gz emd-0510.cif.gz | 6.6 KB | ||
| Others |  emd_0510_half_map_1.map.gz emd_0510_half_map_1.map.gz emd_0510_half_map_2.map.gz emd_0510_half_map_2.map.gz | 132.3 MB 134.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0510 http://ftp.pdbj.org/pub/emdb/structures/EMD-0510 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0510 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0510 | HTTPS FTP |
-Related structure data
| Related structure data |  6ntsMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0510.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0510.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.064 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_0510_msk_1.map emd_0510_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
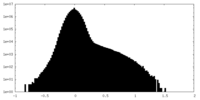
| Density Histograms |
-Half map: #1
| File | emd_0510_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
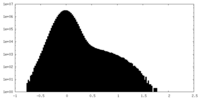
| Density Histograms |
-Half map: #2
| File | emd_0510_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PP2A Aalpha-B56alpha-Calpha holoenzyme in complex with a Small Mo...
| Entire | Name: PP2A Aalpha-B56alpha-Calpha holoenzyme in complex with a Small Molecule Activator of PP2A (SMAP) |
|---|---|
| Components |
|
-Supramolecule #1: PP2A Aalpha-B56alpha-Calpha holoenzyme in complex with a Small Mo...
| Supramolecule | Name: PP2A Aalpha-B56alpha-Calpha holoenzyme in complex with a Small Molecule Activator of PP2A (SMAP) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit...
| Macromolecule | Name: Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 65.378344 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MAAADGDDSL YPIAVLIDEL RNEDVQLRLN SIKKLSTIAL ALGVERTRSE LLPFLTDTIY DEDEVLLALA EQLGTFTTLV GGPEYVHCL LPPLESLATV EETVVRDKAV ESLRAISHEH SPSDLEAHFV PLVKRLAGGD WFTSRTSACG LFSVCYPRVS S AVKAELRQ ...String: MAAADGDDSL YPIAVLIDEL RNEDVQLRLN SIKKLSTIAL ALGVERTRSE LLPFLTDTIY DEDEVLLALA EQLGTFTTLV GGPEYVHCL LPPLESLATV EETVVRDKAV ESLRAISHEH SPSDLEAHFV PLVKRLAGGD WFTSRTSACG LFSVCYPRVS S AVKAELRQ YFRNLCSDDT PMVRRAAASK LGEFAKVLEL DNVKSEIIPM FSNLASDEQD SVRLLAVEAC VNIAQLLPQE DL EALVMPT LRQAAEDKSW RVRYMVADKF TELQKAVGPE ITKTDLVPAF QNLMKDCEAE VRAAASHKVK EFCENLSADC REN VIMSQI LPCIKELVSD ANQHVKSALA SVIMGLSPIL GKDNTIEHLL PLFLAQLKDE CPEVRLNIIS NLDCVNEVIG IRQL SQSLL PAIVELAEDA KWRVRLAIIE YMPLLAGQLG VEFFDEKLNS LCMAWLVDHV YAIREAATSN LKKLVEKFGK EWAHA TIIP KVLAMSGDPN YLHRMTTLFC INVLSEVCGQ DITTKHMLPT VLRMAGDPVA NVRFNVAKSL QKIGPILDNS TLQSEV KPI LEKLTQDQDV DVKYFAQEAL TVLSLA UniProtKB: Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform |
-Macromolecule #2: Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit...
| Macromolecule | Name: Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit alpha isoform type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.266555 KDa |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: MSSSSPPAGA ASAAISASEK VDGFTRKSVR KAQRQKRSQG SSQFRSQGSQ AELHPLPQLK DATSNEQQEL FCQKLQQCCI LFDFMDSVS DLKSKEIKRA TLNELVEYVS TNRGVIVESA YSDIVKMISA NIFRTLPPSD NPDFDPEEDE PTLEASWPHI Q LVYEFFLR ...String: MSSSSPPAGA ASAAISASEK VDGFTRKSVR KAQRQKRSQG SSQFRSQGSQ AELHPLPQLK DATSNEQQEL FCQKLQQCCI LFDFMDSVS DLKSKEIKRA TLNELVEYVS TNRGVIVESA YSDIVKMISA NIFRTLPPSD NPDFDPEEDE PTLEASWPHI Q LVYEFFLR FLESPDFQPS IAKRYIDQKF VQQLLELFDS EDPRERDFLK TVLHRIYGKF LGLRAFIRKQ INNIFLRFIY ET EHFNGVA ELLEILGSII NGFALPLKAE HKQFLMKVLI PMHTAKGLAL FHAQLAYCVV QFLEKDTTLT EPVIRGLLKF WPK TCSQKE VMFLGEIEEI LDVIEPTQFK KIEEPLFKQI SKCVSSSHFQ VAERALYFWN NEYILSLIEE NIDKILPIMF ASLY KISKE HWNPTIVALV YNVLKTLMEM NGKLFDDLTS SYKAERQREK KKELEREELW KKLEELKLKK ALEKQNSAYN MHSIL SNTS AE UniProtKB: Serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit alpha isoform |
-Macromolecule #3: Serine/threonine-protein phosphatase 2A catalytic subunit alpha i...
| Macromolecule | Name: Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: protein-serine/threonine phosphatase |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 35.0215 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MDEKVFTKEL DQWIEQLNEC KQLSESQVKS LCEKAKEILT KESNVQEVRC PVTVCGDVHG QFHDLMELFR IGGKSPDTNY LFMGDYVNR GYYSVETVTL LVALKVRYRE RITILRGNHE SRQITQVYGF YDECLRKYGN ANVWKYFTDL FDYLPLTALV D GQIFCLHG ...String: MDEKVFTKEL DQWIEQLNEC KQLSESQVKS LCEKAKEILT KESNVQEVRC PVTVCGDVHG QFHDLMELFR IGGKSPDTNY LFMGDYVNR GYYSVETVTL LVALKVRYRE RITILRGNHE SRQITQVYGF YDECLRKYGN ANVWKYFTDL FDYLPLTALV D GQIFCLHG GLSPSIDTLD HIRALDRLQE VPHEGPMCDL LWSDPDDRGG WGISPRGAGY TFGQDISETF NHANGLTLVS RA HQLVMEG YNWCHDRNVV TIFSAPNYCY RCGNQAAIME LDDTLKYSFL QFDPAPRRGE PHVTYF(MLL) UniProtKB: Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform |
-Macromolecule #4: N-[(1R,2R,3S)-2-hydroxy-3-(10H-phenoxazin-10-yl)cyclohexyl]-4-(tr...
| Macromolecule | Name: N-[(1R,2R,3S)-2-hydroxy-3-(10H-phenoxazin-10-yl)cyclohexyl]-4-(trifluoromethoxy)benzene-1-sulfonamide type: ligand / ID: 4 / Number of copies: 1 / Formula: L2J |
|---|---|
| Molecular weight | Theoretical: 520.521 Da |
| Chemical component information | 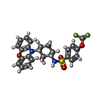 ChemComp-L2J: |
-Macromolecule #5: MANGANESE (II) ION
| Macromolecule | Name: MANGANESE (II) ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: MN |
|---|---|
| Molecular weight | Theoretical: 54.938 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.02 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z
Z Y
Y X
X