[English] 日本語
 Yorodumi
Yorodumi- EMDB-0487: Cryo-EM structure of the TRPM8 ion channel in complex with the me... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0487 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the TRPM8 ion channel in complex with the menthol analog WS-12 and PI(4,5)P2 | |||||||||
 Map data Map data | em-volume_P1 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Ficedula albicollis (Collared flycatcher) Ficedula albicollis (Collared flycatcher) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Yin Y / Le SC / Hsu AL / Borgnia MJ / Yang H / Lee S-Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2019 Journal: Science / Year: 2019Title: Structural basis of cooling agent and lipid sensing by the cold-activated TRPM8 channel. Authors: Ying Yin / Son C Le / Allen L Hsu / Mario J Borgnia / Huanghe Yang / Seok-Yong Lee /  Abstract: Transient receptor potential melastatin member 8 (TRPM8) is a calcium ion (Ca)-permeable cation channel that serves as the primary cold and menthol sensor in humans. Activation of TRPM8 by cooling ...Transient receptor potential melastatin member 8 (TRPM8) is a calcium ion (Ca)-permeable cation channel that serves as the primary cold and menthol sensor in humans. Activation of TRPM8 by cooling compounds relies on allosteric actions of agonist and membrane lipid phosphatidylinositol 4,5-bisphosphate (PIP), but lack of structural information has thus far precluded a mechanistic understanding of ligand and lipid sensing by TRPM8. Using cryo-electron microscopy, we determined the structures of TRPM8 in complex with the synthetic cooling compound icilin, PIP, and Ca, as well as in complex with the menthol analog WS-12 and PIP Our structures reveal the binding sites for cooling agonists and PIP in TRPM8. Notably, PIP binds to TRPM8 in two different modes, which illustrate the mechanism of allosteric coupling between PIP and agonists. This study provides a platform for understanding the molecular mechanism of TRPM8 activation by cooling agents. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0487.map.gz emd_0487.map.gz | 58.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0487-v30.xml emd-0487-v30.xml emd-0487.xml emd-0487.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0487.png emd_0487.png | 288.8 KB | ||
| Others |  emd_0487_half_map_1.map.gz emd_0487_half_map_1.map.gz emd_0487_half_map_2.map.gz emd_0487_half_map_2.map.gz | 45.6 MB 45.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0487 http://ftp.pdbj.org/pub/emdb/structures/EMD-0487 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0487 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0487 | HTTPS FTP |
-Validation report
| Summary document |  emd_0487_validation.pdf.gz emd_0487_validation.pdf.gz | 779.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0487_full_validation.pdf.gz emd_0487_full_validation.pdf.gz | 779.4 KB | Display | |
| Data in XML |  emd_0487_validation.xml.gz emd_0487_validation.xml.gz | 12 KB | Display | |
| Data in CIF |  emd_0487_validation.cif.gz emd_0487_validation.cif.gz | 14.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0487 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0487 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0487 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0487 | HTTPS FTP |
-Related structure data
| Related structure data |  6nr2MC  0488C  0489C  6nr3C  6nr4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0487.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0487.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | em-volume_P1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: half map 1
| File | emd_0487_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_0487_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Transient receptor potential melastatin member 8
| Entire | Name: Transient receptor potential melastatin member 8 |
|---|---|
| Components |
|
-Supramolecule #1: Transient receptor potential melastatin member 8
| Supramolecule | Name: Transient receptor potential melastatin member 8 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Ficedula albicollis (Collared flycatcher) Ficedula albicollis (Collared flycatcher) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Transient receptor potential cation channel subfamily M member 8
| Macromolecule | Name: Transient receptor potential cation channel subfamily M member 8 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ficedula albicollis (Collared flycatcher) Ficedula albicollis (Collared flycatcher) |
| Molecular weight | Theoretical: 130.078453 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MATLFQVSMG SMRHRRNGNF ESSRLLYSSM SRSIDVACSD ADLANFIQEN FKKRECVFFT KDTKSMGNLC KCGYPENQHI EGTQVNTTE KWNYKKH(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) ...String: MATLFQVSMG SMRHRRNGNF ESSRLLYSSM SRSIDVACSD ADLANFIQEN FKKRECVFFT KDTKSMGNLC KCGYPENQHI EGTQVNTTE KWNYKKH(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)K YIRLSCDTDS ETLYDLMTQH WHLKTPNLVI SVTG GAKNF ALKPRMRKIF SRLIYIAQSK GAWIFTGGTH YGLMKYIGEV VRDNTISRSS EENVVAIGIA AWGMISNRET LIRTA DSDG SYLAHYIMDD LKRDPLYCLD NNHTHLLLVD NGTHGHPTIE AKVRTQLEKY ISERVIPESN YGGKIPIVCF AQGGGK ETL KSINVAIKSK IPCVVVEGSG RIADVIASLV EAEGTLASSC VKESLLRFLP RTISRLSEEE TESWIKWIKE VLESPHL LT VIKIEEAGDE IVSNAISFAL YKAFSTNEHD RDNWNGQLKL LLEWNQLDLA SDEIFTNDRN WESADLQDVM FTALVKDR P KFVRLFLENG LNLRKFLTTE VLRELYTNNF SSLVFKNLQI AKNSYNDALL TFVWKMVEDF RRGAKRDDKN SKDEMEIEL SEECPITRHP LQALFIWSVL QNKKELSKVI WEQTRGCTLA ALGASKLLKS MAKVKNDINA AGESEELANE YETRAVELFT ECYSNDEDL AEQLLTYSCE AWGGSNCLEL AVEARDQQFI AQPGVQNFLS KQWYGEISRD TKNWKIILCL FFFPLIGCGF I SFRKKPVE KTKKLFLYYV SFFTSPFVVF SWNVIFYIAF LLLFAYVLLM DFQKEPTALE IILYVLVFIL LCDEVRQWYM NG SKYFSDL WNVMDTLAIF YFIAGIVFRL HSDESSWYSG RVIFCLDYIV FTLRLIHIFT VSRNLGPKII MLQRMMIDVF FFL FLFAVW MVAFGVARQG ILRKNEHRWE WIFRSVIYEP YLAMFGQYPD DIDGTTYNFD HCTFSGNESK PLCVELDANN QPRF PEWIT IPLVCIYMLS TNILLVNLLV AMFGYTVGSV QENNDQVWKF QRFFLVQEYC SRLTIPFPFV IFAYIFMVMR KCFKC CCKK ESKEPSVCCS RNEDNEILAW EAVMKENYLV KINTKASDSS EE(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)KI KSNSLEVLFQ GPDYKDDDDK AHHHHHHHHH H |
-Macromolecule #2: (2S)-1-{[(R)-hydroxy{[(1R,2R,3S,4R,5R,6S)-2,3,6-trihydroxy-4,5-bi...
| Macromolecule | Name: (2S)-1-{[(R)-hydroxy{[(1R,2R,3S,4R,5R,6S)-2,3,6-trihydroxy-4,5-bis(phosphonooxy)cyclohexyl]oxy}phosphoryl]oxy}-3-(octadecanoyloxy)propan-2-yl icosa-5,8,11,14-tetraenoate type: ligand / ID: 2 / Number of copies: 4 / Formula: KXP |
|---|---|
| Molecular weight | Theoretical: 1.047088 KDa |
| Chemical component information | 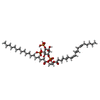 ChemComp-KXP: |
-Macromolecule #3: (1R,2S,5R)-N-(4-methoxyphenyl)-5-methyl-2-(propan-2-yl)cyclohexan...
| Macromolecule | Name: (1R,2S,5R)-N-(4-methoxyphenyl)-5-methyl-2-(propan-2-yl)cyclohexane-1-carboxamide type: ligand / ID: 3 / Number of copies: 4 / Formula: KXS |
|---|---|
| Molecular weight | Theoretical: 289.412 Da |
| Chemical component information | 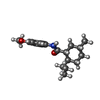 ChemComp-KXS: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 2.0 nm / Pretreatment - Type: GLOW DISCHARGE / Details: 15 mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 4006 / Average exposure time: 60.0 sec. / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.25 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X


















