+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDC29 |
|---|---|
 Sample Sample | Human Rev7 dimer in complex with Rev3 peptide @ 10.6 mg/mL
|
| Function / homology |  Function and homology information Function and homology informationsomatic diversification of immunoglobulins involved in immune response / DNA damage response, signal transduction resulting in transcription /  telomere maintenance in response to DNA damage / negative regulation of transcription regulatory region DNA binding / zeta DNA polymerase complex / positive regulation of isotype switching / positive regulation of extracellular matrix assembly / negative regulation of transcription by competitive promoter binding / negative regulation of cell-cell adhesion mediated by cadherin / negative regulation of epithelial to mesenchymal transition ...somatic diversification of immunoglobulins involved in immune response / DNA damage response, signal transduction resulting in transcription / telomere maintenance in response to DNA damage / negative regulation of transcription regulatory region DNA binding / zeta DNA polymerase complex / positive regulation of isotype switching / positive regulation of extracellular matrix assembly / negative regulation of transcription by competitive promoter binding / negative regulation of cell-cell adhesion mediated by cadherin / negative regulation of epithelial to mesenchymal transition ...somatic diversification of immunoglobulins involved in immune response / DNA damage response, signal transduction resulting in transcription /  telomere maintenance in response to DNA damage / negative regulation of transcription regulatory region DNA binding / zeta DNA polymerase complex / positive regulation of isotype switching / positive regulation of extracellular matrix assembly / negative regulation of transcription by competitive promoter binding / negative regulation of cell-cell adhesion mediated by cadherin / negative regulation of epithelial to mesenchymal transition / JUN kinase binding / negative regulation of ubiquitin protein ligase activity / mitotic spindle assembly checkpoint signaling / positive regulation of double-strand break repair via nonhomologous end joining / negative regulation of double-strand break repair via homologous recombination / error-prone translesion synthesis / positive regulation of epithelial to mesenchymal transition / Translesion synthesis by REV1 / Translesion synthesis by POLK / Translesion synthesis by POLI / actin filament organization / telomere maintenance in response to DNA damage / negative regulation of transcription regulatory region DNA binding / zeta DNA polymerase complex / positive regulation of isotype switching / positive regulation of extracellular matrix assembly / negative regulation of transcription by competitive promoter binding / negative regulation of cell-cell adhesion mediated by cadherin / negative regulation of epithelial to mesenchymal transition / JUN kinase binding / negative regulation of ubiquitin protein ligase activity / mitotic spindle assembly checkpoint signaling / positive regulation of double-strand break repair via nonhomologous end joining / negative regulation of double-strand break repair via homologous recombination / error-prone translesion synthesis / positive regulation of epithelial to mesenchymal transition / Translesion synthesis by REV1 / Translesion synthesis by POLK / Translesion synthesis by POLI / actin filament organization /  regulation of cell growth / double-strand break repair via homologous recombination / negative regulation of canonical Wnt signaling pathway / negative regulation of protein catabolic process / negative regulation of DNA-binding transcription factor activity / DNA-templated DNA replication / spindle / double-strand break repair / positive regulation of peptidyl-serine phosphorylation / site of double-strand break / regulation of cell growth / double-strand break repair via homologous recombination / negative regulation of canonical Wnt signaling pathway / negative regulation of protein catabolic process / negative regulation of DNA-binding transcription factor activity / DNA-templated DNA replication / spindle / double-strand break repair / positive regulation of peptidyl-serine phosphorylation / site of double-strand break /  chromosome / 4 iron, 4 sulfur cluster binding / RNA polymerase II-specific DNA-binding transcription factor binding / chromosome / 4 iron, 4 sulfur cluster binding / RNA polymerase II-specific DNA-binding transcription factor binding /  DNA-directed DNA polymerase / DNA-directed DNA polymerase /  DNA-directed DNA polymerase activity / DNA-directed DNA polymerase activity /  cell division / cell division /  nucleotide binding / nucleotide binding /  chromatin / chromatin /  nucleolus / positive regulation of DNA-templated transcription / negative regulation of transcription by RNA polymerase II / nucleolus / positive regulation of DNA-templated transcription / negative regulation of transcription by RNA polymerase II /  DNA binding / DNA binding /  nucleoplasm / nucleoplasm /  metal ion binding / metal ion binding /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function |
| Biological species |   Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2018 Journal: Proc Natl Acad Sci U S A / Year: 2018Title: Rev7 dimerization is important for assembly and function of the Rev1/Polζ translesion synthesis complex. Authors: Alessandro A Rizzo / Faye-Marie Vassel / Nimrat Chatterjee / Sanjay D'Souza / Yunfeng Li / Bing Hao / Michael T Hemann / Graham C Walker / Dmitry M Korzhnev /  Abstract: The translesion synthesis (TLS) polymerases Polζ and Rev1 form a complex that enables replication of damaged DNA. The Rev7 subunit of Polζ, which is a multifaceted HORMA (Hop1, Rev7, Mad2) protein ...The translesion synthesis (TLS) polymerases Polζ and Rev1 form a complex that enables replication of damaged DNA. The Rev7 subunit of Polζ, which is a multifaceted HORMA (Hop1, Rev7, Mad2) protein with roles in TLS, DNA repair, and cell-cycle control, facilitates assembly of this complex by binding Rev1 and the catalytic subunit of Polζ, Rev3. Rev7 interacts with Rev3 by a mechanism conserved among HORMA proteins, whereby an open-to-closed transition locks the ligand underneath the "safety belt" loop. Dimerization of HORMA proteins promotes binding and release of this ligand, as exemplified by the Rev7 homolog, Mad2. Here, we investigate the dimerization of Rev7 when bound to the two Rev7-binding motifs (RBMs) in Rev3 by combining in vitro analyses of Rev7 structure and interactions with a functional assay in a Rev7 cell line. We demonstrate that Rev7 uses the conventional HORMA dimerization interface both to form a homodimer when tethered by the two RBMs in Rev3 and to heterodimerize with other HORMA domains, Mad2 and p31 Structurally, the Rev7 dimer can bind only one copy of Rev1, revealing an unexpected Rev1/Polζ architecture. In cells, mutation of the Rev7 dimer interface increases sensitivity to DNA damage. These results provide insights into the structure of the Rev1/Polζ TLS assembly and highlight the function of Rev7 homo- and heterodimerization. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDC29 SASDC29 |
|---|
-Related structure data
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
| Model #1625 |  Type: atomic / Radius of dummy atoms: 1.90 A Comment: residues at N- and C-termini that were missing in PDB were built in extended conformation Chi-square value: 25.1295213698  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|
- Sample
Sample
 Sample Sample | Name: Human Rev7 dimer in complex with Rev3 peptide @ 10.6 mg/mL Specimen concentration: 10.6 mg/ml / Entity id: 857 / 858 / 859 |
|---|---|
| Buffer | Name: 20 mM HEPES, 10 mM DTT, 5% glycerol / pH: 8 |
| Entity #857 | Name: hRev7-WT dimer / Type: protein Description: Mitotic spindle assembly checkpoint protein MAD2B Formula weight: 24.391 / Num. of mol.: 2 / Source: Homo sapiens / References: UniProt: Q9UI95 Sequence: GMTTLTRQDL NFGQVVADVL CEFLEVAVHL ILYVREVYPV GIFQKRKKYN VPVQMSCHPE LNQYIQDTLH CVKPLLEKND VEKVVVVILD KEHRPVEKFV FEITQPPLLS ISSDSLLSHV EQLLRAFILK ISVCDAVLDH NPPGCTFTVL VHTREAATRN MEKIQVIKDF ...Sequence: GMTTLTRQDL NFGQVVADVL CEFLEVAVHL ILYVREVYPV GIFQKRKKYN VPVQMSCHPE LNQYIQDTLH CVKPLLEKND VEKVVVVILD KEHRPVEKFV FEITQPPLLS ISSDSLLSHV EQLLRAFILK ISVCDAVLDH NPPGCTFTVL VHTREAATRN MEKIQVIKDF PWILADEQDV HMHDPRLIPL KTMTSDILKM QLYVEERAHK GS |
| Entity #858 | Name: Rev3-RBM2 / Type: protein / Description: DNA polymerase zeta catalytic subunit / Formula weight: 3.273 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: O60673 Sequence: MEDKKIVIMP CKCAPSRQLV QVWLQAKE |
| Entity #859 | Name: Rev3-RBM2 / Type: protein / Description: DNA polymerase zeta catalytic subunit / Formula weight: 3.273 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: O60673 Sequence: MEDKKIVIMP CKCAPSRQLV QVWLQAKE |
-Experimental information
| Beam | Instrument name: Cornell High Energy Synchrotron Source (CHESS) G1 City: Ithaca, NY / 国: USA  / Type of source: X-ray synchrotron / Type of source: X-ray synchrotron Synchrotron / Wavelength: 0.1267 Å Synchrotron / Wavelength: 0.1267 Å | |||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Finger Lakes CCD / Type: CCD | |||||||||||||||||||||||||||||||||||||||
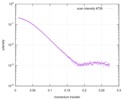
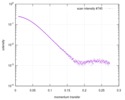
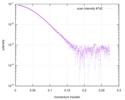
| Scan | Measurement date: May 14, 2016 / Storage temperature: 4 °C / Cell temperature: 4 °C / Exposure time: 1 sec. / Number of frames: 10 / Unit: 1/A /
| |||||||||||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| |||||||||||||||||||||||||||||||||||||||
| Result | Comments: Model was created by mapping the dimer interface by mutagenesis (9 residues identified) and then docking the dimer with HADDOCK and the mutations. SAXS/WAXS was used for cross-validation.
|
 Movie
Movie Controller
Controller