[English] 日本語
 Yorodumi
Yorodumi- SASDAU4: uPAR H47C/N259C (Urokinase plasminogen activator surface receptor... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDAU4 |
|---|---|
 Sample Sample | uPAR H47C/N259C
|
| Function / homology |  Function and homology information Function and homology information urokinase plasminogen activator receptor activity / Attachment of GPI anchor to uPAR / positive regulation of homotypic cell-cell adhesion / negative regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation / urokinase plasminogen activator receptor activity / Attachment of GPI anchor to uPAR / positive regulation of homotypic cell-cell adhesion / negative regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation /  regulation of fibrinolysis / protein complex involved in cell-matrix adhesion / regulation of fibrinolysis / protein complex involved in cell-matrix adhesion /  regulation of proteolysis / serine-type endopeptidase complex ... regulation of proteolysis / serine-type endopeptidase complex ... urokinase plasminogen activator receptor activity / Attachment of GPI anchor to uPAR / positive regulation of homotypic cell-cell adhesion / negative regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation / urokinase plasminogen activator receptor activity / Attachment of GPI anchor to uPAR / positive regulation of homotypic cell-cell adhesion / negative regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway / urokinase plasminogen activator signaling pathway / regulation of plasminogen activation /  regulation of fibrinolysis / protein complex involved in cell-matrix adhesion / regulation of fibrinolysis / protein complex involved in cell-matrix adhesion /  regulation of proteolysis / serine-type endopeptidase complex / Dissolution of Fibrin Clot / positive regulation of epidermal growth factor receptor signaling pathway / regulation of proteolysis / serine-type endopeptidase complex / Dissolution of Fibrin Clot / positive regulation of epidermal growth factor receptor signaling pathway /  extrinsic component of membrane / positive regulation of release of cytochrome c from mitochondria / positive regulation of DNA binding / extrinsic component of membrane / positive regulation of release of cytochrome c from mitochondria / positive regulation of DNA binding /  regulation of cell adhesion / specific granule membrane / negative regulation of intrinsic apoptotic signaling pathway / cell projection / regulation of cell adhesion / specific granule membrane / negative regulation of intrinsic apoptotic signaling pathway / cell projection /  chemotaxis / chemotaxis /  blood coagulation / blood coagulation /  signaling receptor activity / positive regulation of protein phosphorylation / protein domain specific binding / external side of plasma membrane / signaling receptor activity / positive regulation of protein phosphorylation / protein domain specific binding / external side of plasma membrane /  endoplasmic reticulum lumen / endoplasmic reticulum lumen /  signaling receptor binding / signaling receptor binding /  focal adhesion / Neutrophil degranulation / endoplasmic reticulum membrane / negative regulation of apoptotic process / focal adhesion / Neutrophil degranulation / endoplasmic reticulum membrane / negative regulation of apoptotic process /  enzyme binding / enzyme binding /  cell surface / cell surface /  signal transduction / extracellular region / signal transduction / extracellular region /  membrane / membrane /  plasma membrane plasma membraneSimilarity search - Function |
| Biological species |   Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Journal: J Biol Chem / Year: 2012 Journal: J Biol Chem / Year: 2012Title: A flexible multidomain structure drives the function of the urokinase-type plasminogen activator receptor (uPAR). Authors: Haydyn D T Mertens / Magnus Kjaergaard / Simon Mysling / Henrik Gårdsvoll / Thomas J D Jørgensen / Dmitri I Svergun / Michael Ploug /  Abstract: The urokinase-type plasminogen activator receptor (uPAR) provides a rendezvous between proteolytic degradation of the extracellular matrix and integrin-mediated adhesion to vitronectin. These ...The urokinase-type plasminogen activator receptor (uPAR) provides a rendezvous between proteolytic degradation of the extracellular matrix and integrin-mediated adhesion to vitronectin. These processes are, however, tightly linked because the high affinity binding of urokinase regulates the binding of uPAR to matrix-embedded vitronectin. Although crystal structures exist to define the corresponding static bi- and trimolecular receptor complexes, it is evident that the dynamic property of uPAR plays a decisive role in its function. In the present study, we combine small angle x-ray scattering, hydrogen-deuterium exchange, and surface plasmon resonance to develop a structural model describing the allosteric regulation of uPAR. We show that the flexibility of its N-terminal domain provides the key for understanding this allosteric mechanism. Importantly, our model has direct implications for understanding uPAR-assisted cell adhesion and migration as well as for translational research, including targeted intervention therapy and non-invasive tumor imaging in vivo. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDAU4 SASDAU4 |
|---|
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
| Model #78 |  Type: dummy / Software: dammif / Radius of dummy atoms: 1.90 A / Symmetry  : P1 / Chi-square value: 2.217121 : P1 / Chi-square value: 2.217121 Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|---|
| Model #80 |  Type: mix / Software: coral / Radius of dummy atoms: 1.90 A / Symmetry  : P1 / Chi-square value: 1.787569 : P1 / Chi-square value: 1.787569 Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
- Sample
Sample
 Sample Sample | Name: uPAR H47C/N259C / Sample MW: 36.98 kDa / Specimen concentration: 1.50-4.40 / Concentration method: A280 |
|---|---|
| Buffer | Name: Sodium Phosphate / Concentration: 25.00 mM / pH: 7.2 / Composition: Glycerol 5.000 %, NaSO4 50.000 mM |
| Entity #66 | Name: uPAR Urokinase receptor / Type: protein Urokinase receptor / Type: proteinDescription: Urokinase plasminogen activator surface receptor  Urokinase receptor Urokinase receptorFormula weight: 36.98 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: Q03405 Sequence: MGHPPLLPLL LLLHTCVPAS WGLRCMQCKT NGDCRVEECA LGQDLCRTTI VRLWEEGEEL ELVEKSCTHS EKTNRTLSYR TGLKITSLTE VVCGLDLCNQ GNSGRAVTYS RSRYLECISC GSSDMSCERG RHQSLQCRSP EEQCLDVVTH WIQEGEEGRP KDDRHLRGCG ...Sequence: MGHPPLLPLL LLLHTCVPAS WGLRCMQCKT NGDCRVEECA LGQDLCRTTI VRLWEEGEEL ELVEKSCTHS EKTNRTLSYR TGLKITSLTE VVCGLDLCNQ GNSGRAVTYS RSRYLECISC GSSDMSCERG RHQSLQCRSP EEQCLDVVTH WIQEGEEGRP KDDRHLRGCG YLPGCPGSNG FHNNDTFHFL KCCNTTKCNE GPILELENLP QNGRQCYSCK GNSTHGCSSE ETFLIDCRGP MNQCLVATGT HEPKNQSYMV RGCATASMCQ HAHLGDAFSM NHIDVSCCTK SGCNHPDLDV QYRSGAAPQP GPAHLSLTIT LLMTARLWGG TLLWT |
-Experimental information
| Beam | Instrument name:  DORIS III X33 DORIS III X33  / City: Hamburg / 国: Germany / City: Hamburg / 国: Germany  / Shape / Shape : 0.6 / Type of source: X-ray synchrotron : 0.6 / Type of source: X-ray synchrotron Synchrotron / Wavelength: 0.15 Å / Dist. spec. to detc.: 2.7 mm Synchrotron / Wavelength: 0.15 Å / Dist. spec. to detc.: 2.7 mm | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 1M-W / Pixsize x: 0.172 mm | ||||||||||||||||||||||||||||||
| Scan |
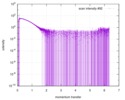
| ||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller