[English] 日本語
 Yorodumi
Yorodumi- PDB-7sqj: Cryo-EM structure of the seam subunits of the enteropathogenic E.... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7sqj | ||||||
|---|---|---|---|---|---|---|---|
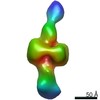
| Title | Cryo-EM structure of the seam subunits of the enteropathogenic E. coli O127:H6 flagellar filament | ||||||
 Components Components | Flagellin | ||||||
 Keywords Keywords |  STRUCTURAL PROTEIN / Bacteria flagellar filament / STRUCTURAL PROTEIN / Bacteria flagellar filament /  motility / flagellar polymorphism motility / flagellar polymorphism | ||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum / structural molecule activity / extracellular region Similarity search - Function | ||||||
| Biological species |   Escherichia coli O127:H6 (bacteria) Escherichia coli O127:H6 (bacteria) | ||||||
| Method |  ELECTRON MICROSCOPY / ELECTRON MICROSCOPY /  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 6.3 Å cryo EM / Resolution: 6.3 Å | ||||||
 Authors Authors | Kreutzberger, M.A.B. / Chatterjee, S. / Frankel, G. / Egelman, E.H. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Flagellin outer domain dimerization modulates motility in pathogenic and soil bacteria from viscous environments. Authors: Mark A B Kreutzberger / Richard C Sobe / Amber B Sauder / Sharanya Chatterjee / Alejandro Peña / Fengbin Wang / Jorge A Giron / Volker Kiessling / Tiago R D Costa / Vincent P Conticello / ...Authors: Mark A B Kreutzberger / Richard C Sobe / Amber B Sauder / Sharanya Chatterjee / Alejandro Peña / Fengbin Wang / Jorge A Giron / Volker Kiessling / Tiago R D Costa / Vincent P Conticello / Gad Frankel / Melissa M Kendall / Birgit E Scharf / Edward H Egelman /   Abstract: Flagellar filaments function as the propellers of the bacterial flagellum and their supercoiling is key to motility. The outer domains on the surface of the filament are non-critical for motility in ...Flagellar filaments function as the propellers of the bacterial flagellum and their supercoiling is key to motility. The outer domains on the surface of the filament are non-critical for motility in many bacteria and their structures and functions are not conserved. Here, we show the atomic cryo-electron microscopy structures for flagellar filaments from enterohemorrhagic Escherichia coli O157:H7, enteropathogenic E. coli O127:H6, Achromobacter, and Sinorhizobium meliloti, where the outer domains dimerize or tetramerize to form either a sheath or a screw-like surface. These dimers are formed by 180° rotations of half of the outer domains. The outer domain sheath (ODS) plays a role in bacterial motility by stabilizing an intermediate waveform and prolonging the tumbling of E. coli cells. Bacteria with these ODS and screw-like flagellar filaments are commonly found in soil and human intestinal environments of relatively high viscosity suggesting a role for the dimerization in these environments. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7sqj.cif.gz 7sqj.cif.gz | 173.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7sqj.ent.gz pdb7sqj.ent.gz | 146.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7sqj.json.gz 7sqj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/sq/7sqj https://data.pdbj.org/pub/pdb/validation_reports/sq/7sqj ftp://data.pdbj.org/pub/pdb/validation_reports/sq/7sqj ftp://data.pdbj.org/pub/pdb/validation_reports/sq/7sqj | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  25386MC  7sn4C  7sn7C  7sn9C  7sqdC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein |  Mass: 56342.008 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Escherichia coli O127:H6 (bacteria) / Gene: fliC, BQ9544_2041 / Production host: Escherichia coli O127:H6 (bacteria) / Gene: fliC, BQ9544_2041 / Production host:   Escherichia coli O127:H6 (bacteria) / References: UniProt: B7USU2 Escherichia coli O127:H6 (bacteria) / References: UniProt: B7USU2 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  ELECTRON MICROSCOPY ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: FILAMENT / 3D reconstruction method:  single particle reconstruction single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Bacterial flagellar filament / Type: COMPLEX / Entity ID: all / Source: MULTIPLE SOURCES |
|---|---|
| Source (natural) | Organism:   Escherichia coli O127:H6 (bacteria) Escherichia coli O127:H6 (bacteria) |
| Buffer solution | pH: 7.2 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied : NO / Vitrification applied : NO / Vitrification applied : YES : YES |
Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: TFS KRIOS |
| Electron gun | Electron source : :  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Electron dose: 50 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) |
- Processing
Processing
| Software | Name: UCSF ChimeraX / Version: 1.2/v9 / Classification: model building / URL: https://www.rbvi.ucsf.edu/chimerax/ / Os: macOS / Type: package |
|---|---|
CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
3D reconstruction | Resolution: 6.3 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 60359 / Symmetry type: POINT |
 Movie
Movie Controller
Controller













 PDBj
PDBj