[English] 日本語
 Yorodumi
Yorodumi- EMDB-34544: Structure of the dimeric Xenopus tropical acid-sensitive outwardl... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 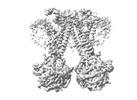 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the dimeric Xenopus tropical acid-sensitive outwardly rectifying channel ASOR trimer bound with tRNA (closed state) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Chloride Channel tRNA Complex /  MEMBRANE PROTEIN MEMBRANE PROTEIN | |||||||||
| Function / homology | pH-gated chloride channel activity / TMEM206 protein / TMEM206 protein family / chloride transport /  chloride channel complex / chloride channel complex /  cell surface / cell surface /  plasma membrane / Proton-activated chloride channel plasma membrane / Proton-activated chloride channel Function and homology information Function and homology information | |||||||||
| Biological species |  Xenopus tropicalis (tropical clawed frog) / Xenopus tropicalis (tropical clawed frog) /   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.81 Å cryo EM / Resolution: 3.81 Å | |||||||||
 Authors Authors | Chi P / Wang X / Li J / Li K / Zhang Y / Geng J / Wu J / Deng D | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of the dimeric Xenopus tropical acid-sensitive outwardly rectifying channel ASOR trimer bound with tRNA (closed state) Authors: Chi P / Wang X / Li J / Li K / Zhang Y / Geng J / Wu J / Deng D | |||||||||
| History |
|
- Structure visualization
Structure visualization
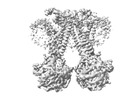
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34544.map.gz emd_34544.map.gz | 230.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34544-v30.xml emd-34544-v30.xml emd-34544.xml emd-34544.xml | 14.1 KB 14.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34544.png emd_34544.png | 78.7 KB | ||
| Filedesc metadata |  emd-34544.cif.gz emd-34544.cif.gz | 5.7 KB | ||
| Others |  emd_34544_half_map_1.map.gz emd_34544_half_map_1.map.gz emd_34544_half_map_2.map.gz emd_34544_half_map_2.map.gz | 226.3 MB 226.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34544 http://ftp.pdbj.org/pub/emdb/structures/EMD-34544 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34544 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34544 | HTTPS FTP |
-Related structure data
| Related structure data |  8h8eMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_34544.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34544.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34544_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34544_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of Xenopus tropical acid-sensitive outwardly rectifying c...
| Entire | Name: Complex of Xenopus tropical acid-sensitive outwardly rectifying channel ASOR trimer and tRNA in digitonin |
|---|---|
| Components |
|
-Supramolecule #1: Complex of Xenopus tropical acid-sensitive outwardly rectifying c...
| Supramolecule | Name: Complex of Xenopus tropical acid-sensitive outwardly rectifying channel ASOR trimer and tRNA in digitonin type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Xenopus tropicalis (tropical clawed frog) Xenopus tropicalis (tropical clawed frog) |
-Macromolecule #1: Proton-activated chloride channel
| Macromolecule | Name: Proton-activated chloride channel / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Xenopus tropicalis (tropical clawed frog) Xenopus tropicalis (tropical clawed frog) |
| Molecular weight | Theoretical: 41.031789 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MEAIRKELSR SYQELNEEAE PVAIDPEEAE DEEKEQEEAA SAVAPDRDSD RSSPPVRFSR TCLKNFFSVL LILVYLLLMG VAVFLVYQT ITDFRDKLKH PVMSVSYKEV NMYDAPGIAL YPGKARLLSC EHHWYDHIPP LKDPGQPGEN TCVTQDISYI D PYTNKTMK ...String: MEAIRKELSR SYQELNEEAE PVAIDPEEAE DEEKEQEEAA SAVAPDRDSD RSSPPVRFSR TCLKNFFSVL LILVYLLLMG VAVFLVYQT ITDFRDKLKH PVMSVSYKEV NMYDAPGIAL YPGKARLLSC EHHWYDHIPP LKDPGQPGEN TCVTQDISYI D PYTNKTMK HALIVQGPRD VRRRELVFLQ FHLNETKQDF SAIDYLLFSS YEAFLKSHDQ VKFMQDCESS FSSWKFSGGF RT WVKMSLV KTKEEDGSQS VEFRQETSVV NFIDRRETPD KGDQLFFVVF EWKDPYIQEI QDIITANPWS MIALLCSVFL VLF KAADFA KLSVKWMIKV RRRHLKKRAR ELNHIS UniProtKB: Proton-activated chloride channel |
-Macromolecule #2: tRNA (75-MER)of Spodoptera frugiperda
| Macromolecule | Name: tRNA (75-MER)of Spodoptera frugiperda / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Molecular weight | Theoretical: 24.108287 KDa |
| Sequence | String: UCUUCGGUAG UAUAGUGGUC AGUAUCCCCG CCUGUCACGC GGGAGACCGG GGUUCGAUUC CCCGCCGGAG AGCCA |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.1 µm Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.1 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 57.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.81 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 182807 |
 Movie
Movie Controller
Controller


 Z
Z Y
Y X
X