+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-2785 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Architecture of the RNA polymerase II-Mediator core transcription initiation complex | |||||||||
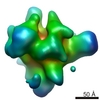
 マップデータ マップデータ | cITC | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード |  transcription (転写 (生物学)) / transcription (転写 (生物学)) /  transcription initiation (転写 (生物学)) / transcription initiation (転写 (生物学)) /  RNA polymerase II (RNAポリメラーゼII) / RNA polymerase II (RNAポリメラーゼII) /  General Transcription Factors (基本転写因子) General Transcription Factors (基本転写因子) | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報RNA polymerase II complex recruiting activity / TFIIA-class transcription factor complex binding / RNA polymerase III transcription regulatory region sequence-specific DNA binding / transcription factor TFIIIB complex / RNA polymerase III preinitiation complex assembly / RNA polymerase I general transcription initiation factor binding / transcription open complex formation at RNA polymerase II promoter / regulation of transcription by RNA polymerase III / transcriptional start site selection at RNA polymerase II promoter / RPB4-RPB7 complex ...RNA polymerase II complex recruiting activity / TFIIA-class transcription factor complex binding / RNA polymerase III transcription regulatory region sequence-specific DNA binding / transcription factor TFIIIB complex / RNA polymerase III preinitiation complex assembly / RNA polymerase I general transcription initiation factor binding / transcription open complex formation at RNA polymerase II promoter / regulation of transcription by RNA polymerase III / transcriptional start site selection at RNA polymerase II promoter / RPB4-RPB7 complex / positive regulation of transcription regulatory region DNA binding / transcription factor TFIIF complex / transcription factor TFIIA complex / RNA polymerase I preinitiation complex assembly / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay /  転写開始前複合体 / DNA binding, bending / RNA Polymerase I Transcription Initiation / Processing of Capped Intron-Containing Pre-mRNA / RNA Polymerase III Transcription Initiation From Type 2 Promoter / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / mRNA Capping / RNA polymerase II transcribes snRNA genes / termination of RNA polymerase II transcription / TP53 Regulates Transcription of DNA Repair Genes / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA-templated transcription / termination of RNA polymerase III transcription / RNA Polymerase II Pre-transcription Events / : / 転写開始前複合体 / DNA binding, bending / RNA Polymerase I Transcription Initiation / Processing of Capped Intron-Containing Pre-mRNA / RNA Polymerase III Transcription Initiation From Type 2 Promoter / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / mRNA Capping / RNA polymerase II transcribes snRNA genes / termination of RNA polymerase II transcription / TP53 Regulates Transcription of DNA Repair Genes / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA-templated transcription / termination of RNA polymerase III transcription / RNA Polymerase II Pre-transcription Events / : /  : / Formation of TC-NER Pre-Incision Complex / termination of RNA polymerase I transcription / RNA Polymerase I Promoter Escape / : / Formation of TC-NER Pre-Incision Complex / termination of RNA polymerase I transcription / RNA Polymerase I Promoter Escape /  transcription initiation at RNA polymerase III promoter / RNA polymerase II complex binding / nucleolar large rRNA transcription by RNA polymerase I / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / Gap-filling DNA repair synthesis and ligation in TC-NER / transcription initiation at RNA polymerase III promoter / RNA polymerase II complex binding / nucleolar large rRNA transcription by RNA polymerase I / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / Gap-filling DNA repair synthesis and ligation in TC-NER /  transcription initiation at RNA polymerase I promoter / Estrogen-dependent gene expression / transcription elongation by RNA polymerase I / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / Dual incision in TC-NER / transcription factor TFIID complex / RNA polymerase II general transcription initiation factor activity / tRNA transcription by RNA polymerase III / transcription initiation at RNA polymerase I promoter / Estrogen-dependent gene expression / transcription elongation by RNA polymerase I / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / Dual incision in TC-NER / transcription factor TFIID complex / RNA polymerase II general transcription initiation factor activity / tRNA transcription by RNA polymerase III /  RNA polymerase III activity / transcription by RNA polymerase I / transcription by RNA polymerase III / RNA polymerase III activity / transcription by RNA polymerase I / transcription by RNA polymerase III /  RNA polymerase I activity / RNA polymerase II core promoter sequence-specific DNA binding / positive regulation of translational initiation / RNA polymerase I activity / RNA polymerase II core promoter sequence-specific DNA binding / positive regulation of translational initiation /  RNA polymerase I complex / RNA polymerase I complex /  RNA polymerase III complex / transcription-coupled nucleotide-excision repair / RNA polymerase III complex / transcription-coupled nucleotide-excision repair /  RNA polymerase II activity / RNA polymerase II activity /  DNA修復 / DNA修復 /  RNA polymerase II, core complex / RNA polymerase II preinitiation complex assembly / RNA polymerase II, core complex / RNA polymerase II preinitiation complex assembly /  translation initiation factor binding / TBP-class protein binding / DNA-templated transcription initiation / transcription elongation by RNA polymerase II / translation initiation factor binding / TBP-class protein binding / DNA-templated transcription initiation / transcription elongation by RNA polymerase II /  transcription initiation at RNA polymerase II promoter / positive regulation of transcription elongation by RNA polymerase II / transcription initiation at RNA polymerase II promoter / positive regulation of transcription elongation by RNA polymerase II /  P-body / P-body /  ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity /  ポリメラーゼ / cytoplasmic stress granule / ポリメラーゼ / cytoplasmic stress granule /  ペルオキシソーム / disordered domain specific binding / ペルオキシソーム / disordered domain specific binding /  リボソーム生合成 / リボソーム生合成 /  single-stranded DNA binding / DNA-binding transcription factor binding / RNA polymerase II-specific DNA-binding transcription factor binding / single-stranded DNA binding / DNA-binding transcription factor binding / RNA polymerase II-specific DNA-binding transcription factor binding /  transcription regulator complex / transcription by RNA polymerase II / transcription regulator complex / transcription by RNA polymerase II /  nucleic acid binding / nucleic acid binding /  single-stranded RNA binding / single-stranded RNA binding /  protein dimerization activity / protein dimerization activity /  nucleotide binding / negative regulation of DNA-templated transcription / nucleotide binding / negative regulation of DNA-templated transcription /  mRNA binding / mRNA binding /  chromatin binding / regulation of DNA-templated transcription / chromatin binding / regulation of DNA-templated transcription /  核小体 / positive regulation of transcription by RNA polymerase II / protein-containing complex / 核小体 / positive regulation of transcription by RNA polymerase II / protein-containing complex /  ミトコンドリア / ミトコンドリア /  DNA binding / zinc ion binding / DNA binding / zinc ion binding /  核質 / 核質 /  metal ion binding / metal ion binding /  細胞核 細胞核類似検索 - 分子機能 | |||||||||
| 生物種 |   Saccharomyces cerevisiae (パン酵母) / Saccharomyces cerevisiae (パン酵母) /   Saccharomyces mikatae (酵母) / synthetic construct (人工物) Saccharomyces mikatae (酵母) / synthetic construct (人工物) | |||||||||
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 7.8 Å クライオ電子顕微鏡法 / 解像度: 7.8 Å | |||||||||
 データ登録者 データ登録者 | Plaschka C / Lariviere L / Wenzeck L / Hemann M / Tegunov D / Petrotchenko EV / Borchers CH / Baumeister W / Herzog F / Villa E / Cramer P | |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2015 ジャーナル: Nature / 年: 2015タイトル: Architecture of the RNA polymerase II-Mediator core initiation complex. 著者: C Plaschka / L Larivière / L Wenzeck / M Seizl / M Hemann / D Tegunov / E V Petrotchenko / C H Borchers / W Baumeister / F Herzog / E Villa / P Cramer /    要旨: The conserved co-activator complex Mediator enables regulated transcription initiation by RNA polymerase (Pol) II. Here we reconstitute an active 15-subunit core Mediator (cMed) comprising all ...The conserved co-activator complex Mediator enables regulated transcription initiation by RNA polymerase (Pol) II. Here we reconstitute an active 15-subunit core Mediator (cMed) comprising all essential Mediator subunits from Saccharomyces cerevisiae. The cryo-electron microscopic structure of cMed bound to a core initiation complex was determined at 9.7 Å resolution. cMed binds Pol II around the Rpb4-Rpb7 stalk near the carboxy-terminal domain (CTD). The Mediator head module binds the Pol II dock and the TFIIB ribbon and stabilizes the initiation complex. The Mediator middle module extends to the Pol II foot with a 'plank' that may influence polymerase conformation. The Mediator subunit Med14 forms a 'beam' between the head and middle modules and connects to the tail module that is predicted to bind transcription activators located on upstream DNA. The Mediator 'arm' and 'hook' domains contribute to a 'cradle' that may position the CTD and TFIIH kinase to stimulate Pol II phosphorylation. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_2785.map.gz emd_2785.map.gz | 77.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-2785-v30.xml emd-2785-v30.xml emd-2785.xml emd-2785.xml | 25.6 KB 25.6 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_2785.png emd_2785.png | 100.4 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2785 http://ftp.pdbj.org/pub/emdb/structures/EMD-2785 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2785 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2785 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_2785.map.gz / 形式: CCP4 / 大きさ: 81.8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_2785.map.gz / 形式: CCP4 / 大きさ: 81.8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | cITC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : cITC
+超分子 #1000: cITC
+分子 #1: DNA-DIRECTED RNA POLYMERASE II SUBUNIT RPB1
+分子 #2: DNA-DIRECTED RNA POLYMERASE II SUBUNIT RPB2
+分子 #3: DNA-DIRECTED RNA POLYMERASE II SUBUNIT RPB3
+分子 #4: DNA-DIRECTED RNA POLYMERASE II SUBUNIT RPB4
+分子 #5: DNA-DIRECTED RNA POLYMERASES I, II, AND III SUBUNIT RPABC 1
+分子 #6: DNA-DIRECTED RNA POLYMERASES I, II, AND III SUBUNIT RPABC 2
+分子 #7: DNA-DIRECTED RNA POLYMERASE II SUBUNIT RPB7
+分子 #8: DNA-DIRECTED RNA POLYMERASES I, II, AND III SUBUNIT RPABC 3
+分子 #9: DNA-DIRECTED RNA POLYMERASE II SUBUNIT RPB9
+分子 #10: DNA-DIRECTED RNA POLYMERASES I, II, AND III SUBUNIT RPABC 5
+分子 #11: DNA-DIRECTED RNA POLYMERASE II SUBUNIT RPB11
+分子 #12: DNA-DIRECTED RNA POLYMERASES I, II, AND III SUBUNIT RPABC 4
+分子 #13: TRANSCRIPTION INITIATION FACTOR IIB
+分子 #14: TATA-box-binding protein
+分子 #15: Transcription initiation factor IIF subunit alpha
+分子 #16: Transcription initiation factor IIF subunit beta
+分子 #17: template DNA
+分子 #18: nontemplate DNA
+分子 #19: RNA
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.15 mg/mL |
|---|---|
| 緩衝液 | pH: 7.5 詳細: 25 mM HEPES-KOH pH 7.5, 180 mM Potassium acetate, 5 % Glycerol, 5 mM DTT |
| 凍結 | 凍結剤: ETHANE / 装置: OTHER 手法: Grids were glow-discharged for 20 s before deposition of 4 microliters sample and incubated for 30 s. Grids were washed twice with 4 microliters distilled water, blotted, and then vitrified. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 倍率(補正後): 37169 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy / Cs: 2 mm / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 1.0 µm / 倍率(公称値): 37000 Bright-field microscopy / Cs: 2 mm / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 1.0 µm / 倍率(公称値): 37000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER |
| 撮影 | カテゴリ: CCD / フィルム・検出器のモデル: GATAN K2 (4k x 4k) / 実像数: 2972 / 平均電子線量: 25 e/Å2 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
| CTF補正 | 詳細: Each particle |
|---|---|
| 最終 再構成 | 想定した対称性 - 点群: C1 (非対称) / アルゴリズム: OTHER / 解像度のタイプ: BY AUTHOR / 解像度: 7.8 Å / 解像度の算出法: OTHER / ソフトウェア - 名称: RELION, 1.2 詳細: The density was filtered according to local resolution and further a temperature factor of minus 340 A2 was applied. 使用した粒子像数: 4439 |
| 詳細 | Particles were picked using EMAN2 and were processed using RELION 1.2. |
 ムービー
ムービー コントローラー
コントローラー