+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
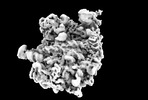
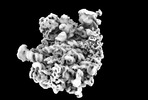
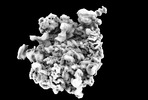
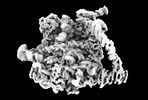
| タイトル | Yeast RQC complex in state E | |||||||||
 マップデータ マップデータ | Yeast RQC complex in state E | |||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報positive regulation of cytoplasmic translational elongation through polyproline stretches / Hypusine synthesis from eIF5A-lysine / CAT tailing /  translational frameshifting / positive regulation of translational termination / RQC complex / positive regulation of translational elongation / Antigen processing: Ubiquitination & Proteasome degradation / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosome-associated ubiquitin-dependent protein catabolic process ...positive regulation of cytoplasmic translational elongation through polyproline stretches / Hypusine synthesis from eIF5A-lysine / CAT tailing / translational frameshifting / positive regulation of translational termination / RQC complex / positive regulation of translational elongation / Antigen processing: Ubiquitination & Proteasome degradation / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosome-associated ubiquitin-dependent protein catabolic process ...positive regulation of cytoplasmic translational elongation through polyproline stretches / Hypusine synthesis from eIF5A-lysine / CAT tailing /  translational frameshifting / positive regulation of translational termination / RQC complex / positive regulation of translational elongation / Antigen processing: Ubiquitination & Proteasome degradation / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosome-associated ubiquitin-dependent protein catabolic process / pre-mRNA 5'-splice site binding / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / translational elongation / response to cycloheximide / maturation of 5.8S rRNA / SRP-dependent cotranslational protein targeting to membrane / translational frameshifting / positive regulation of translational termination / RQC complex / positive regulation of translational elongation / Antigen processing: Ubiquitination & Proteasome degradation / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosome-associated ubiquitin-dependent protein catabolic process / pre-mRNA 5'-splice site binding / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / translational elongation / response to cycloheximide / maturation of 5.8S rRNA / SRP-dependent cotranslational protein targeting to membrane /  ribosomal large subunit binding / GTP hydrolysis and joining of the 60S ribosomal subunit / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / negative regulation of mRNA splicing, via spliceosome / preribosome, large subunit precursor / positive regulation of translational initiation / L13a-mediated translational silencing of Ceruloplasmin expression / ribosomal large subunit binding / GTP hydrolysis and joining of the 60S ribosomal subunit / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / negative regulation of mRNA splicing, via spliceosome / preribosome, large subunit precursor / positive regulation of translational initiation / L13a-mediated translational silencing of Ceruloplasmin expression /  ribosomal large subunit export from nucleus / subtelomeric heterochromatin formation / regulation of translational fidelity / ribosomal large subunit export from nucleus / subtelomeric heterochromatin formation / regulation of translational fidelity /  ribosomal subunit export from nucleus / ribosomal subunit export from nucleus /  translation elongation factor activity / 90S preribosome / cytosolic ribosome / rescue of stalled ribosome / translation elongation factor activity / 90S preribosome / cytosolic ribosome / rescue of stalled ribosome /  translation initiation factor activity / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / assembly of large subunit precursor of preribosome / proteasomal protein catabolic process / translation initiation factor activity / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / assembly of large subunit precursor of preribosome / proteasomal protein catabolic process /  ribosomal large subunit biogenesis / cytosolic ribosome assembly / ribosomal large subunit biogenesis / cytosolic ribosome assembly /  オートファジー / RING-type E3 ubiquitin transferase / protein catabolic process / オートファジー / RING-type E3 ubiquitin transferase / protein catabolic process /  ribosomal large subunit assembly / modification-dependent protein catabolic process / ribosomal large subunit assembly / modification-dependent protein catabolic process /  protein tag activity / rRNA processing / ubiquitin-protein transferase activity / protein tag activity / rRNA processing / ubiquitin-protein transferase activity /  ubiquitin protein ligase activity / ubiquitin protein ligase activity /  リボソーム生合成 / リボソーム生合成 /  ribosome binding / chromatin organization / large ribosomal subunit rRNA binding / ribosome binding / chromatin organization / large ribosomal subunit rRNA binding /  5S rRNA binding / ubiquitin-dependent protein catabolic process / cytoplasmic translation / cytosolic large ribosomal subunit / 5S rRNA binding / ubiquitin-dependent protein catabolic process / cytoplasmic translation / cytosolic large ribosomal subunit /  chromosome, telomeric region / negative regulation of translation / chromosome, telomeric region / negative regulation of translation /  rRNA binding / rRNA binding /  リボソーム / protein ubiquitination / structural constituent of ribosome / リボソーム / protein ubiquitination / structural constituent of ribosome /  ribonucleoprotein complex / ribonucleoprotein complex /  翻訳 (生物学) / response to antibiotic / 翻訳 (生物学) / response to antibiotic /  mRNA binding / mRNA binding /  ubiquitin protein ligase binding / ubiquitin protein ligase binding /  核小体 / perinuclear region of cytoplasm / 核小体 / perinuclear region of cytoplasm /  ミトコンドリア / ミトコンドリア /  RNA binding / zinc ion binding / RNA binding / zinc ion binding /  metal ion binding / metal ion binding /  細胞核 / 細胞核 /  細胞質基質 / 細胞質基質 /  細胞質 細胞質類似検索 - 分子機能 | |||||||||
| 生物種 |   Saccharomyces cerevisiae (パン酵母) / Saccharomyces cerevisiae (パン酵母) /   baker's yeast (パン酵母) baker's yeast (パン酵母) | |||||||||
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 2.7 Å クライオ電子顕微鏡法 / 解像度: 2.7 Å | |||||||||
 データ登録者 データ登録者 | Tesina P / Buschauer R / Beckmann R | |||||||||
| 資金援助 |  ドイツ, 1件 ドイツ, 1件
| |||||||||
 引用 引用 |  ジャーナル: Mol Cell / 年: 2023 ジャーナル: Mol Cell / 年: 2023タイトル: Molecular basis of eIF5A-dependent CAT tailing in eukaryotic ribosome-associated quality control. 著者: Petr Tesina / Shuhei Ebine / Robert Buschauer / Matthias Thoms / Yoshitaka Matsuo / Toshifumi Inada / Roland Beckmann /   要旨: Ribosome-associated quality control (RQC) is a conserved process degrading potentially toxic truncated nascent peptides whose malfunction underlies neurodegeneration and proteostasis decline in aging. ...Ribosome-associated quality control (RQC) is a conserved process degrading potentially toxic truncated nascent peptides whose malfunction underlies neurodegeneration and proteostasis decline in aging. During RQC, dissociation of stalled ribosomes is followed by elongation of the nascent peptide with alanine and threonine residues, driven by Rqc2 independently of mRNA, the small ribosomal subunit and guanosine triphosphate (GTP)-hydrolyzing factors. The resulting CAT tails (carboxy-terminal tails) and ubiquitination by Ltn1 mark nascent peptides for proteasomal degradation. Here we present ten cryogenic electron microscopy (cryo-EM) structures, revealing the mechanistic basis of individual steps of the CAT tailing cycle covering initiation, decoding, peptidyl transfer, and tRNA translocation. We discovered eIF5A as a crucial eukaryotic RQC factor enabling peptidyl transfer. Moreover, we observed dynamic behavior of RQC factors and tRNAs allowing for processivity of the CAT tailing cycle without additional energy input. Together, these results elucidate key differences as well as common principles between CAT tailing and canonical translation. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_15424.map.gz emd_15424.map.gz | 20.8 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-15424-v30.xml emd-15424-v30.xml emd-15424.xml emd-15424.xml | 69 KB 69 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_15424_fsc.xml emd_15424_fsc.xml | 15.6 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_15424.png emd_15424.png | 60.8 KB | ||
| その他 |  emd_15424_half_map_1.map.gz emd_15424_half_map_1.map.gz emd_15424_half_map_2.map.gz emd_15424_half_map_2.map.gz | 322.3 MB 322.3 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15424 http://ftp.pdbj.org/pub/emdb/structures/EMD-15424 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15424 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15424 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8aguMC  8aafC  8agtC  8agvC  8agwC  8agxC  8agzC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_15424.map.gz / 形式: CCP4 / 大きさ: 347.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_15424.map.gz / 形式: CCP4 / 大きさ: 347.6 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Yeast RQC complex in state E | ||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.059 Å | ||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: Yeast RQC complex in state E, half map A
| ファイル | emd_15424_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Yeast RQC complex in state E, half map A | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Yeast RQC complex in state E, half map B
| ファイル | emd_15424_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Yeast RQC complex in state E, half map B | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : Yeast RQC complex in state E
+超分子 #1: Yeast RQC complex in state E
+分子 #1: 60S ribosomal protein L15-A
+分子 #2: 60S ribosomal protein L16-A
+分子 #3: 60S ribosomal protein L17-A
+分子 #4: 60S ribosomal protein L18-A
+分子 #5: 60S ribosomal protein L19-A
+分子 #6: 60S ribosomal protein L20-A
+分子 #7: 60S ribosomal protein L21-A
+分子 #8: 60S ribosomal protein L22-A
+分子 #9: 60S ribosomal protein L23-A
+分子 #10: 60S ribosomal protein L24-A
+分子 #11: 60S ribosomal protein L25
+分子 #12: 60S ribosomal protein L26-A
+分子 #13: 60S ribosomal protein L27-A
+分子 #14: 60S ribosomal protein L28
+分子 #15: 60S ribosomal protein L29
+分子 #16: 60S ribosomal protein L30
+分子 #17: 60S ribosomal protein L31-A
+分子 #18: 60S ribosomal protein L32
+分子 #19: 60S ribosomal protein L33-A
+分子 #20: 60S ribosomal protein L34-A
+分子 #21: 60S ribosomal protein L35-A
+分子 #22: 60S ribosomal protein L36-A
+分子 #23: 60S ribosomal protein L37-A
+分子 #24: BJ4_G0032190.mRNA.1.CDS.1
+分子 #25: 60S ribosomal protein L39
+分子 #26: Ubiquitin-60S ribosomal protein L40
+分子 #27: 60S ribosomal protein L42-A
+分子 #28: 60S ribosomal protein L43-A
+分子 #29: RPL41A isoform 1
+分子 #33: 60S ribosomal protein L2-A
+分子 #34: 60S ribosomal protein L3
+分子 #35: BJ4_G0008850.mRNA.1.CDS.1
+分子 #36: 60S ribosomal protein L5
+分子 #37: 60S ribosomal protein L6-B
+分子 #38: 60S ribosomal protein L7-A
+分子 #39: 60S ribosomal protein L8-A
+分子 #40: 60S ribosomal protein L9-A
+分子 #41: 60S ribosomal protein L10
+分子 #42: BJ4_G0027750.mRNA.1.CDS.1
+分子 #43: 60S ribosomal protein L13-A
+分子 #44: 60S ribosomal protein L14-A
+分子 #45: RQC2 isoform 1
+分子 #46: E3 ubiquitin-protein ligase listerin
+分子 #47: Eukaryotic translation initiation factor 6
+分子 #48: Eukaryotic translation initiation factor 5A-1
+分子 #49: 60S ribosomal protein L1-A
+分子 #51: 60S ribosomal protein L12-B
+分子 #52: 60S acidic ribosomal protein P0
+分子 #53: CAT-tailed nascent peptide
+分子 #30: 25S rRNA
+分子 #31: 5S rRNA
+分子 #32: 5.8S rRNA
+分子 #50: P-site tRNA
+分子 #54: MAGNESIUM ION
+分子 #55: ZINC ION
+分子 #56: SPERMIDINE
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 6.8 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy / 最大 デフォーカス(公称値): 4.0 µm / 最小 デフォーカス(公称値): 0.4 µm Bright-field microscopy / 最大 デフォーカス(公称値): 4.0 µm / 最小 デフォーカス(公称値): 0.4 µm |
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 平均電子線量: 46.0 e/Å2 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー






















 Z
Z Y
Y X
X