[English] 日本語
 Yorodumi
Yorodumi- EMDB-14240: Early transcription elongation state of influenza B polymerase ba... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Early transcription elongation state of influenza B polymerase backtracked due to double incoproation of nucleotide analogue T1106 | |||||||||
 Map data Map data | LocScale filtered | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity /  cap snatching / 7-methylguanosine mRNA capping / viral transcription / host cell mitochondrion / cap snatching / 7-methylguanosine mRNA capping / viral transcription / host cell mitochondrion /  virion component / virion component /  endonuclease activity / host cell cytoplasm / endonuclease activity / host cell cytoplasm /  Hydrolases; Acting on ester bonds / Hydrolases; Acting on ester bonds /  RNA-directed RNA polymerase ...symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / RNA-directed RNA polymerase ...symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity /  cap snatching / 7-methylguanosine mRNA capping / viral transcription / host cell mitochondrion / cap snatching / 7-methylguanosine mRNA capping / viral transcription / host cell mitochondrion /  virion component / virion component /  endonuclease activity / host cell cytoplasm / endonuclease activity / host cell cytoplasm /  Hydrolases; Acting on ester bonds / Hydrolases; Acting on ester bonds /  RNA-directed RNA polymerase / viral RNA genome replication / RNA-directed RNA polymerase / viral RNA genome replication /  RNA-dependent RNA polymerase activity / RNA-dependent RNA polymerase activity /  nucleotide binding / DNA-templated transcription / host cell nucleus / nucleotide binding / DNA-templated transcription / host cell nucleus /  RNA binding / RNA binding /  metal ion binding metal ion bindingSimilarity search - Function | |||||||||
| Biological species |   Influenza B virus (B/Memphis/13/2003) / Influenza B virus (B/Memphis/13/2003) /   Influenza B virus Influenza B virus | |||||||||
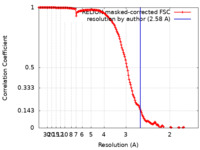
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.58 Å cryo EM / Resolution: 2.58 Å | |||||||||
 Authors Authors | Cusack S / Kouba T | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2023 Journal: Cell Rep / Year: 2023Title: Direct observation of backtracking by influenza A and B polymerases upon consecutive incorporation of the nucleoside analog T1106. Authors: Tomas Kouba / Anna Dubankova / Petra Drncova / Elisa Donati / Pietro Vidossich / Valentina Speranzini / Alex Pflug / Johanna Huchting / Chris Meier / Marco De Vivo / Stephen Cusack /    Abstract: The antiviral pseudo-base T705 and its de-fluoro analog T1106 mimic adenine or guanine and can be competitively incorporated into nascent RNA by viral RNA-dependent RNA polymerases. Although ...The antiviral pseudo-base T705 and its de-fluoro analog T1106 mimic adenine or guanine and can be competitively incorporated into nascent RNA by viral RNA-dependent RNA polymerases. Although dispersed, single pseudo-base incorporation is mutagenic, consecutive incorporation causes polymerase stalling and chain termination. Using a template encoding single and then consecutive T1106 incorporation four nucleotides later, we obtained a cryogenic electron microscopy structure of stalled influenza A/H7N9 polymerase. This shows that the entire product-template duplex backtracks by 5 nt, bringing the singly incorporated T1106 to the +1 position, where it forms an unexpected T1106:U wobble base pair. Similar structures show that influenza B polymerase also backtracks after consecutive T1106 incorporation, regardless of whether prior single incorporation has occurred. These results give insight into the unusual mechanism of chain termination by pyrazinecarboxamide base analogs. Consecutive incorporation destabilizes the proximal end of the product-template duplex, promoting irreversible backtracking to a more energetically favorable overall configuration. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14240.map.gz emd_14240.map.gz | 159.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14240-v30.xml emd-14240-v30.xml emd-14240.xml emd-14240.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14240_fsc.xml emd_14240_fsc.xml | 15.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_14240.png emd_14240.png | 122.9 KB | ||
| Others |  emd_14240_additional_1.map.gz emd_14240_additional_1.map.gz emd_14240_half_map_1.map.gz emd_14240_half_map_1.map.gz emd_14240_half_map_2.map.gz emd_14240_half_map_2.map.gz | 304.5 MB 259.9 MB 259.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14240 http://ftp.pdbj.org/pub/emdb/structures/EMD-14240 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14240 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14240 | HTTPS FTP |
-Related structure data
| Related structure data |  7r1fMC  7qtlC  7r0eC  7zplC  7zpmC  8bdrC  8be0C  8bekC  8bf5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14240.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14240.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LocScale filtered | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: RELION post-processed
| File | emd_14240_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | RELION post-processed | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_14240_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_14240_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : B/Mem-BACKTRACKED
+Supramolecule #1: B/Mem-BACKTRACKED
+Macromolecule #1: Polymerase acidic protein
+Macromolecule #2: RNA-directed RNA polymerase catalytic subunit
+Macromolecule #3: Polymerase basic protein 2
+Macromolecule #4: 5' vRNA
+Macromolecule #5: 3' vRNA
+Macromolecule #6: mRNA
+Macromolecule #7: MAGNESIUM ION
+Macromolecule #8: P1-7-METHYLGUANOSINE-P3-ADENOSINE-5',5'-TRIPHOSPHATE
+Macromolecule #9: water
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.5 µm Bright-field microscopy / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.5 µm |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 33.4 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-7r1f: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)