[English] 日本語
 Yorodumi
Yorodumi- EMDB-14076: Cryo-EM structure of human full-length synaptic alpha1beta3gamma2... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human full-length synaptic alpha1beta3gamma2 GABA(A)R in complex with Ro15-4513 and megabody Mb38 | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationGABA receptor activation / GABA-gated chloride ion channel activity / cellular response to histamine /  inhibitory synapse assembly / inhibitory synapse assembly /  GABA-A receptor activity / GABA-A receptor activity /  GABA-A receptor complex / GABA-A receptor complex /  neurotransmitter receptor activity / neurotransmitter receptor activity /  synaptic transmission, GABAergic / gamma-aminobutyric acid signaling pathway / chloride transport ...GABA receptor activation / GABA-gated chloride ion channel activity / cellular response to histamine / synaptic transmission, GABAergic / gamma-aminobutyric acid signaling pathway / chloride transport ...GABA receptor activation / GABA-gated chloride ion channel activity / cellular response to histamine /  inhibitory synapse assembly / inhibitory synapse assembly /  GABA-A receptor activity / GABA-A receptor activity /  GABA-A receptor complex / GABA-A receptor complex /  neurotransmitter receptor activity / neurotransmitter receptor activity /  synaptic transmission, GABAergic / gamma-aminobutyric acid signaling pathway / chloride transport / synaptic transmission, GABAergic / gamma-aminobutyric acid signaling pathway / chloride transport /  chloride channel activity / roof of mouth development / Signaling by ERBB4 / extracellular ligand-gated monoatomic ion channel activity / chloride channel activity / roof of mouth development / Signaling by ERBB4 / extracellular ligand-gated monoatomic ion channel activity /  chloride channel complex / chloride transmembrane transport / cytoplasmic vesicle membrane / chloride channel complex / chloride transmembrane transport / cytoplasmic vesicle membrane /  postsynaptic membrane / neuron projection / postsynaptic membrane / neuron projection /  synapse / synapse /  signal transduction / identical protein binding / signal transduction / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | ||||||||||||||||||
| Biological species |   Homo sapiens (human) / Homo sapiens (human) /   Lama glama (llama) Lama glama (llama) | ||||||||||||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.7 Å cryo EM / Resolution: 2.7 Å | ||||||||||||||||||
 Authors Authors | Sente A / Desai R / Naydenova K / Malinauskas T / Jounaidi Y / Miehling J / Zhou X / Masiulis S / Hardwick SW / Chirgadze DY ...Sente A / Desai R / Naydenova K / Malinauskas T / Jounaidi Y / Miehling J / Zhou X / Masiulis S / Hardwick SW / Chirgadze DY / Miller KW / Aricescu AR | ||||||||||||||||||
| Funding support |  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Differential assembly diversifies GABA receptor structures and signalling. Authors: Andrija Sente / Rooma Desai / Katerina Naydenova / Tomas Malinauskas / Youssef Jounaidi / Jonas Miehling / Xiaojuan Zhou / Simonas Masiulis / Steven W Hardwick / Dimitri Y Chirgadze / Keith ...Authors: Andrija Sente / Rooma Desai / Katerina Naydenova / Tomas Malinauskas / Youssef Jounaidi / Jonas Miehling / Xiaojuan Zhou / Simonas Masiulis / Steven W Hardwick / Dimitri Y Chirgadze / Keith W Miller / A Radu Aricescu /    Abstract: Type A γ-aminobutyric acid receptors (GABARs) are pentameric ligand-gated chloride channels that mediate fast inhibitory signalling in neural circuits and can be modulated by essential medicines ...Type A γ-aminobutyric acid receptors (GABARs) are pentameric ligand-gated chloride channels that mediate fast inhibitory signalling in neural circuits and can be modulated by essential medicines including general anaesthetics and benzodiazepines. Human GABAR subunits are encoded by 19 paralogous genes that can, in theory, give rise to 495,235 receptor types. However, the principles that govern the formation of pentamers, the permutational landscape of receptors that may emerge from a subunit set and the effect that this has on GABAergic signalling remain largely unknown. Here we use cryogenic electron microscopy to determine the structures of extrasynaptic GABARs assembled from α4, β3 and δ subunits, and their counterparts incorporating γ2 instead of δ subunits. In each case, we identified two receptor subtypes with distinct stoichiometries and arrangements, all four differing from those previously observed for synaptic, α1-containing receptors. This, in turn, affects receptor responses to physiological and synthetic modulators by creating or eliminating ligand-binding sites at subunit interfaces. We provide structural and functional evidence that selected GABAR arrangements can act as coincidence detectors, simultaneously responding to two neurotransmitters: GABA and histamine. Using assembly simulations and single-cell RNA sequencing data, we calculated the upper bounds for receptor diversity in recombinant systems and in vivo. We propose that differential assembly is a pervasive mechanism for regulating the physiology and pharmacology of GABARs. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14076.map.gz emd_14076.map.gz | 4.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14076-v30.xml emd-14076-v30.xml emd-14076.xml emd-14076.xml | 21 KB 21 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14076.png emd_14076.png | 56.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14076 http://ftp.pdbj.org/pub/emdb/structures/EMD-14076 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14076 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14076 | HTTPS FTP |
-Related structure data
| Related structure data |  7qneMC  7qn5C  7qn6C  7qn7C  7qn8C  7qn9C  7qnaC  7qnbC  7qncC  7qndC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-10912 (Title: Cryo-EM micrographs of GABA(A)Rs purified from cells expressing human full-length alpha1, beta3 and gamma subunits, in presence of Ro15-4513 and megabody Mb38 EMPIAR-10912 (Title: Cryo-EM micrographs of GABA(A)Rs purified from cells expressing human full-length alpha1, beta3 and gamma subunits, in presence of Ro15-4513 and megabody Mb38Data size: 1.4 TB Data #1: Unaligned multi-frame micrographs of GABA(A)Rs purified from cells expressing human full-length alpha1, beta3 and gamma subunits, in presence of Ro15-4513 and megabody Mb38 [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14076.map.gz / Format: CCP4 / Size: 93 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14076.map.gz / Format: CCP4 / Size: 93 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.896 Å | ||||||||||||||||||||||||||||||||||||
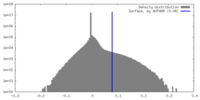
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : Human full-length synaptic alpha1beta3gamma2 GABA(A)R in complex ...
+Supramolecule #1: Human full-length synaptic alpha1beta3gamma2 GABA(A)R in complex ...
+Macromolecule #1: GABA(A) receptor subunit alpha-1
+Macromolecule #2: Gamma-aminobutyric acid receptor subunit beta-3
+Macromolecule #3: GABA(A) receptor subunit gamma-2
+Macromolecule #4: Megabody Mb38
+Macromolecule #9: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(o...
+Macromolecule #10: N-OCTANE
+Macromolecule #11: DECANE
+Macromolecule #12: CHLORIDE ION
+Macromolecule #13: 2-acetamido-2-deoxy-beta-D-glucopyranose
+Macromolecule #14: ethyl 8-[(azanylidene-$l^{4}-azanylidene)amino]-5-methyl-6-oxidan...
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 287 K / Instrument: LEICA PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 130000 Bright-field microscopy / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 130000 |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: CTFFIND (ver. 4.1.13) |
|---|---|
| Startup model | Type of model: OTHER / Details: AB INITIO MODEL |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
| Final 3D classification | Software - Name: RELION (ver. 3.1) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3.1) |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.7 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number images used: 119901 |
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7qne: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)