+Search query
-Structure paper
| Title | Structural basis of odorant recognition by a human odorant receptor. |
|---|---|
| Journal, issue, pages | Nature, Vol. 615, Issue 7953, Page 742-749, Year 2023 |
| Publish date | Mar 15, 2023 |
 Authors Authors | Christian B Billesbølle / Claire A de March / Wijnand J C van der Velden / Ning Ma / Jeevan Tewari / Claudia Llinas Del Torrent / Linus Li / Bryan Faust / Nagarajan Vaidehi / Hiroaki Matsunami / Aashish Manglik /    |
| PubMed Abstract | Our sense of smell enables us to navigate a vast space of chemically diverse odour molecules. This task is accomplished by the combinatorial activation of approximately 400 odorant G protein-coupled ...Our sense of smell enables us to navigate a vast space of chemically diverse odour molecules. This task is accomplished by the combinatorial activation of approximately 400 odorant G protein-coupled receptors encoded in the human genome. How odorants are recognized by odorant receptors remains unclear. Here we provide mechanistic insight into how an odorant binds to a human odorant receptor. Using cryo-electron microscopy, we determined the structure of the active human odorant receptor OR51E2 bound to the fatty acid propionate. Propionate is bound within an occluded pocket in OR51E2 and makes specific contacts critical to receptor activation. Mutation of the odorant-binding pocket in OR51E2 alters the recognition spectrum for fatty acids of varying chain length, suggesting that odorant selectivity is controlled by tight packing interactions between an odorant and an odorant receptor. Molecular dynamics simulations demonstrate that propionate-induced conformational changes in extracellular loop 3 activate OR51E2. Together, our studies provide a high-resolution view of chemical recognition of an odorant by a vertebrate odorant receptor, providing insight into how this large family of G protein-coupled receptors enables our olfactory sense. |
 External links External links |  Nature / Nature /  PubMed:36922591 / PubMed:36922591 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.1 - 3.2 Å |
| Structure data | EMDB-28896, PDB-8f76:  EMDB-28900: Propionate bound to human olfactory receptor OR51E2 in complex with miniGs399 (transmembrane domain) |
| Chemicals | 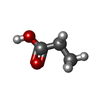 ChemComp-PPI: |
| Source |
|
 Keywords Keywords |  MEMBRANE PROTEIN / MEMBRANE PROTEIN /  Human olfactory receptor / Human olfactory receptor /  G protein-coupled receptor / odorant-binding / G protein-coupled receptor / odorant-binding /  sensory transduction sensory transduction |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers







