+Search query
-Structure paper
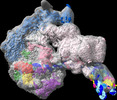
| Title | Yeast PIC-Mediator structure with RNA polymerase II C-terminal domain. |
|---|---|
| Journal, issue, pages | Proc Natl Acad Sci U S A, Vol. 120, Issue 15, Page e2220542120, Year 2023 |
| Publish date | Apr 11, 2023 |
 Authors Authors | Sandra Schilbach / Haibo Wang / Christian Dienemann / Patrick Cramer /  |
| PubMed Abstract | For transcription initiation, RNA polymerase II (Pol II) forms a preinitiation complex (PIC) that associates with the general coactivator Mediator. Whereas atomic models of the human PIC-Mediator ...For transcription initiation, RNA polymerase II (Pol II) forms a preinitiation complex (PIC) that associates with the general coactivator Mediator. Whereas atomic models of the human PIC-Mediator structure have been reported, structures for its yeast counterpart remain incomplete. Here, we present an atomic model for the yeast PIC with core Mediator, including the Mediator middle module that was previously poorly resolved and including subunit Med1 that was previously lacking. We observe three peptide regions containing eleven of the 26 heptapeptide repeats of the flexible C-terminal repeat domain (CTD) of Pol II. Two of these CTD regions bind between the Mediator head and middle modules and form defined CTD-Mediator interactions. CTD peptide 1 binds between the Med6 shoulder and Med31 knob domains, whereas CTD peptide 2 forms additional contacts with Med4. The third CTD region (peptide 3) binds in the Mediator cradle and associates with the Mediator hook. Comparisons with the human PIC-Mediator structure show that the central region in peptide 1 is similar and forms conserved contacts with Mediator, whereas peptides 2 and 3 exhibit distinct structures and Mediator interactions. |
 External links External links |  Proc Natl Acad Sci U S A / Proc Natl Acad Sci U S A /  PubMed:37014863 / PubMed:37014863 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.0 - 3.6 Å |
| Structure data | EMDB-16610, PDB-8cen: EMDB-16611, PDB-8ceo: |
| Chemicals |  ChemComp-SF4:  ChemComp-ZN:  ChemComp-MG: |
| Source |
|
 Keywords Keywords |  TRANSCRIPTION / PIC / Mediator / TF / TRANSCRIPTION / PIC / Mediator / TF /  nucleosome nucleosome |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers