+Search query
-Structure paper
| Title | A structural basis for prion strain diversity. |
|---|---|
| Journal, issue, pages | Nat Chem Biol, Vol. 19, Issue 5, Page 607-613, Year 2023 |
| Publish date | Jan 16, 2023 |
 Authors Authors | Szymon W Manka / Adam Wenborn / Jemma Betts / Susan Joiner / Helen R Saibil / John Collinge / Jonathan D F Wadsworth /  |
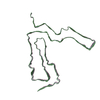
| PubMed Abstract | Recent cryogenic electron microscopy (cryo-EM) studies of infectious, ex vivo, prion fibrils from hamster 263K and mouse RML prion strains revealed a similar, parallel in-register intermolecular β- ...Recent cryogenic electron microscopy (cryo-EM) studies of infectious, ex vivo, prion fibrils from hamster 263K and mouse RML prion strains revealed a similar, parallel in-register intermolecular β-sheet (PIRIBS) amyloid architecture. Rungs of the fibrils are composed of individual prion protein (PrP) monomers that fold to create distinct N-terminal and C-terminal lobes. However, disparity in the hamster/mouse PrP sequence precludes understanding of how divergent prion strains emerge from an identical PrP substrate. In this study, we determined the near-atomic resolution cryo-EM structure of infectious, ex vivo mouse prion fibrils from the ME7 prion strain and compared this with the RML fibril structure. This structural comparison of two biologically distinct mouse-adapted prion strains suggests defined folding subdomains of PrP rungs and the way in which they are interrelated, providing a structural definition of intra-species prion strain-specific conformations. |
 External links External links |  Nat Chem Biol / Nat Chem Biol /  PubMed:36646960 / PubMed:36646960 /  PubMed Central PubMed Central |
| Methods | EM (helical sym.) |
| Resolution | 2.6 Å |
| Structure data | EMDB-15043, PDB-8a00: |
| Source |
|
 Keywords Keywords | PROTEIN FIBRIL / Prion |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers