+Search query
-Structure paper
| Title | HIV-1 CD4-binding site germline antibody-Env structures inform vaccine design. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 13, Issue 1, Page 6123, Year 2022 |
| Publish date | Oct 17, 2022 |
 Authors Authors | Kim-Marie A Dam / Christopher O Barnes / Harry B Gristick / Till Schoofs / Priyanthi N P Gnanapragasam / Michel C Nussenzweig / Pamela J Bjorkman /    |
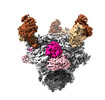
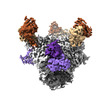
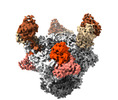
| PubMed Abstract | BG24, a VRC01-class broadly neutralizing antibody (bNAb) against HIV-1 Env with relatively few somatic hypermutations (SHMs), represents a promising target for vaccine strategies to elicit CD4- ...BG24, a VRC01-class broadly neutralizing antibody (bNAb) against HIV-1 Env with relatively few somatic hypermutations (SHMs), represents a promising target for vaccine strategies to elicit CD4-binding site (CD4bs) bNAbs. To understand how SHMs correlate with BG24 neutralization of HIV-1, we report 4.1 Å and 3.4 Å single-particle cryo-EM structures of two inferred germline (iGL) BG24 precursors complexed with engineered Env-based immunogens lacking CD4bs N-glycans. Structures reveal critical Env contacts by BG24 and identify antibody light chain structural features that impede Env recognition. In addition, biochemical data and cryo-EM structures of BG24 variants bound to Envs with CD4bs glycans present provide insights into N-glycan accommodation, including structural modes of light chain adaptations in the presence of the N276 glycan. Together, these findings reveal Env regions critical for germline antibody recognition and potential sites to alter in immunogen design. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:36253376 / PubMed:36253376 /  PubMed Central PubMed Central |
| Methods | EM (single particle) / X-ray diffraction |
| Resolution | 1.4 - 6.4 Å |
| Structure data | EMDB-26490, PDB-7ugn:  EMDB-26491: Cryo-EM structure of BG24 inferred germline Fabs with germline CDR3s and 10-1074 Fabs in complex with HIV-1 Env immunogen BG505-SOSIPv4.1-GT1 - Class 2 EMDB-26492, PDB-7ugo: EMDB-26493, PDB-7ugp:  EMDB-26494: Cryo-EM map of BG24 Fabs with an inferred germline light chain and 10-1074 Fabs in complex with HIV-1 Env immunogen BG505-SOSIPv4.1-GT1 containing the N276 gp120 glycan- Class 2  EMDB-26495: Cryo-EM map of BG24 Fabs with an inferred germline light chain and 10-1074 Fabs in complex with HIV-1 Env immunogen BG505-SOSIPv4.1-GT1 containing the N276 gp120 glycan- Class 3 EMDB-26496, PDB-7ugq:  PDB-7ugm: |
| Chemicals |  ChemComp-HOH:  ChemComp-NAG:  ChemComp-MAN: |
| Source |
|
 Keywords Keywords |  IMMUNE SYSTEM / IMMUNE SYSTEM /  Fab / IgG / BG24 / anti-HIV antibody / bNAb / Fab / IgG / BG24 / anti-HIV antibody / bNAb /  VIRAL PROTEIN/IMMUNE SYSTEM / HIV-1 Env / VIRAL PROTEIN/IMMUNE SYSTEM / HIV-1 Env /  neutralizing antibody / CD4 binding site antibody / neutralizing antibody / CD4 binding site antibody /  VIRAL PROTEIN / VIRAL PROTEIN /  Germline / precursor / Germline / precursor /  immunogen / immunogen /  VIRAL PROTEIN-IMMUNE SYSTEM complex VIRAL PROTEIN-IMMUNE SYSTEM complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers