+Search query
-Structure paper
| Title | Mechanisms of loading and release of the 9-1-1 checkpoint clamp. |
|---|---|
| Journal, issue, pages | Nat Struct Mol Biol, Vol. 29, Issue 4, Page 369-375, Year 2022 |
| Publish date | Mar 21, 2022 |
 Authors Authors | Juan C Castaneda / Marina Schrecker / Dirk Remus / Richard K Hite /  |
| PubMed Abstract | Single-stranded or double-stranded DNA junctions with recessed 5' ends serve as loading sites for the checkpoint clamp, 9-1-1, which mediates activation of the apical checkpoint kinase, ATR. However, ...Single-stranded or double-stranded DNA junctions with recessed 5' ends serve as loading sites for the checkpoint clamp, 9-1-1, which mediates activation of the apical checkpoint kinase, ATR. However, the basis for 9-1-1's recruitment to 5' junctions is unclear. Here, we present structures of the yeast checkpoint clamp loader, Rad24-replication factor C (RFC), in complex with 9-1-1 and a 5' junction and in a post-ATP-hydrolysis state. Unexpectedly, 9-1-1 adopts both closed and planar open states in the presence of Rad24-RFC and DNA. Moreover, Rad24-RFC associates with the DNA junction in the opposite orientation of processivity clamp loaders with Rad24 exclusively coordinating the double-stranded region. ATP hydrolysis stimulates conformational changes in Rad24-RFC, leading to disengagement of DNA-loaded 9-1-1. Together, these structures explain 9-1-1's recruitment to 5' junctions and reveal new principles of sliding clamp loading. |
 External links External links |  Nat Struct Mol Biol / Nat Struct Mol Biol /  PubMed:35314831 / PubMed:35314831 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.2 - 3.9 Å |
| Structure data | EMDB-25422, PDB-7st9: EMDB-25423, PDB-7stb:  EMDB-25424: Mec3 focus refinement of Rad24-RFC:9-1-1:DNA in the open state  EMDB-25425: Rad24-Rfc4 focus refinement of Rad24-RFC-ADP EMDB-25426, PDB-7ste: |
| Chemicals |  ChemComp-AGS:  ChemComp-MG:  ChemComp-GLU: 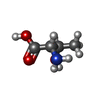 ChemComp-THR:  ChemComp-ADP:  ChemComp-HOH:  ChemComp-ATP: |
| Source |
|
 Keywords Keywords | REPLICATION/DNA /  DNA damage / DNA damage /  DNA replication / DNA sliding clamp / DNA replication / DNA sliding clamp /  REPLICATION / REPLICATION-DNA complex REPLICATION / REPLICATION-DNA complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers










