+Search query
-Structure paper
| Title | De novo design of monomeric helical bundles for pH-controlled membrane lysis. |
|---|---|
| Journal, issue, pages | Protein Sci, Vol. 32, Issue 11, Page e4769, Year 2023 |
| Publish date | Oct 30, 2023 |
 Authors Authors | Nicolas Goldbach / Issa Benna / Basile I M Wicky / Jacob T Croft / Lauren Carter / Asim K Bera / Hannah Nguyen / Alex Kang / Banumathi Sankaran / Erin C Yang / Kelly K Lee / David Baker /   |
| PubMed Abstract | Targeted intracellular delivery via receptor-mediated endocytosis requires the delivered cargo to escape the endosome to prevent lysosomal degradation. This can in principle be achieved by membrane ...Targeted intracellular delivery via receptor-mediated endocytosis requires the delivered cargo to escape the endosome to prevent lysosomal degradation. This can in principle be achieved by membrane lysis tightly restricted to endosomal membranes upon internalization to avoid general membrane insertion and lysis. Here, we describe the design of small monomeric proteins with buried histidine containing pH-responsive hydrogen bond networks and membrane permeating amphipathic helices. Of the 30 designs that were experimentally tested, all expressed in Escherichia coli, 13 were monomeric with the expected secondary structure, and 4 designs disrupted artificial liposomes in a pH-dependent manner. Mutational analysis showed that the buried histidine hydrogen bond networks mediate pH-responsiveness and control lysis of model membranes within a very narrow range of pH (6.0-5.5) with almost no lysis occurring at neutral pH. These tightly controlled lytic monomers could help mediate endosomal escape in designed targeted delivery platforms. |
 External links External links |  Protein Sci / Protein Sci /  PubMed:37632837 / PubMed:37632837 /  PubMed Central PubMed Central |
| Methods | EM (tomography) / X-ray diffraction |
| Resolution | 2.3 Å |
| Structure data |  EMDB-40185: Representative cryo-electron tomogram of pRLB-540 mixed with liposomes at pH 8.0 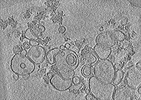 EMDB-40186: Representative cryo-electron tomogram of pRLB-540 mixed with liposomes (acidified to pH 5.5, plunge frozen after 20 s) 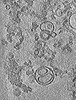 EMDB-40187: Representative cryo-electron tomogram of pRLB-540 mixed with liposomes, acidified to pH 5.5 and plunge frozen after 60s incubation.  PDB-8gl3: |
| Chemicals |  ChemComp-HOH: |
| Source |
|
 Keywords Keywords | DE NOVO PROTEIN / protein design / pH-responsive / membrane lysis |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



