+Search query
-Structure paper
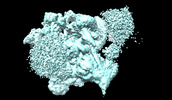
| Title | Structural basis of U12-type intron engagement by the fully assembled human minor spliceosome. |
|---|---|
| Journal, issue, pages | Science, Vol. 383, Issue 6688, Page 1245-1252, Year 2024 |
| Publish date | Mar 15, 2024 |
 Authors Authors | Rui Bai / Meng Yuan / Pu Zhang / Ting Luo / Yigong Shi / Ruixue Wan /  |
| PubMed Abstract | The minor spliceosome, which is responsible for the splicing of U12-type introns, comprises five small nuclear RNAs (snRNAs), of which only one is shared with the major spliceosome. In this work, we ...The minor spliceosome, which is responsible for the splicing of U12-type introns, comprises five small nuclear RNAs (snRNAs), of which only one is shared with the major spliceosome. In this work, we report the 3.3-angstrom cryo-electron microscopy structure of the fully assembled human minor spliceosome pre-B complex. The atomic model includes U11 small nuclear ribonucleoprotein (snRNP), U12 snRNP, and U4atac/U6atac.U5 tri-snRNP. U11 snRNA is recognized by five U11-specific proteins (20K, 25K, 35K, 48K, and 59K) and the heptameric Sm ring. The 3' half of the 5'-splice site forms a duplex with U11 snRNA; the 5' half is recognized by U11-35K, U11-48K, and U11 snRNA. Two proteins, CENATAC and DIM2/TXNL4B, specifically associate with the minor tri-snRNP. A structural analysis uncovered how two conformationally similar tri-snRNPs are differentiated by the minor and major prespliceosomes for assembly. |
 External links External links |  Science / Science /  PubMed:38484052 PubMed:38484052 |
| Methods | EM (single particle) |
| Resolution | 3.13 - 4.66 Å |
| Structure data | EMDB-38993, PDB-8y6o:  EMDB-38999: Focused refinement map of BRR2 region of the human minor pre-B complex  EMDB-39000: focused refinement map for the U11 snRNP in the human minor pre-B complex  EMDB-39005: Focused refinement map of U5 snRNP of the human minor pre-B complex  EMDB-39006: Focused refinement map of U5 Sm ring region of the human minor pre-B complex  EMDB-39007: Focused refinement map for U4atac/U6atac region of the human minor pre-B complex EMDB-39013, PDB-8y7e: |
| Chemicals |  ChemComp-MG:  ChemComp-IHP:  ChemComp-GTP:  ChemComp-ZN: |
| Source |
|
 Keywords Keywords | SPLICING/RNA /  spliceosome / minor spliceosome / U11 snRNP / U12 snRNP / U4atac / U6atac / CENATAC / TXNL4B / spliceosome / minor spliceosome / U11 snRNP / U12 snRNP / U4atac / U6atac / CENATAC / TXNL4B /  U11 snRNA / minor tri-snRNP / U12-type intron / SPLICING-RNA complex U11 snRNA / minor tri-snRNP / U12-type intron / SPLICING-RNA complex |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers