+Search query
-Structure paper
| Title | Structure, adsorption to host, and infection mechanism of virulent lactococcal phage p2. |
|---|---|
| Journal, issue, pages | J Virol, Vol. 87, Issue 22, Page 12302-12312, Year 2013 |
| Publish date | Sep 11, 2013 |
 Authors Authors | Cecilia Bebeacua / Denise Tremblay / Carine Farenc / Marie-Pierre Chapot-Chartier / Irina Sadovskaya / Marin van Heel / David Veesler / Sylvain Moineau / Christian Cambillau /  |
| PubMed Abstract | Lactococcal siphophages from the 936 and P335 groups infect the Gram-positive bacterium Lactococcus lactis using receptor binding proteins (RBPs) attached to their baseplate, a large multiprotein ...Lactococcal siphophages from the 936 and P335 groups infect the Gram-positive bacterium Lactococcus lactis using receptor binding proteins (RBPs) attached to their baseplate, a large multiprotein complex at the distal part of the tail. We have previously reported the crystal and electron microscopy (EM) structures of the baseplates of phages p2 (936 group) and TP901-1 (P335 group) as well as the full EM structure of the TP901-1 virion. Here, we report the complete EM structure of siphophage p2, including its capsid, connector complex, tail, and baseplate. Furthermore, we show that the p2 tail is characterized by the presence of protruding decorations, which are related to adhesins and are likely contributed by the major tail protein C-terminal domains. This feature is reminiscent of the tail of Escherichia coli phage λ and Bacillus subtilis phage SPP1 and might point to a common mechanism for establishing initial interactions with their bacterial hosts. Comparative analyses showed that the architecture of the phage p2 baseplate differs largely from that of lactococcal phage TP901-1. We quantified the interaction of its RBP with the saccharidic receptor and determined that specificity is due to lower k(off) values of the RBP/saccharidic dissociation. Taken together, these results suggest that the infection of L. lactis strains by phage p2 is a multistep process that involves reversible attachment, followed by baseplate activation, specific attachment of the RBPs to the saccharidic receptor, and DNA ejection. |
 External links External links |  J Virol / J Virol /  PubMed:24027307 / PubMed:24027307 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 13.0 - 22.0 Å |
| Structure data |  EMDB-2459:  EMDB-2462:  EMDB-2463: 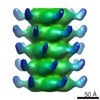 EMDB-2464: |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



