+Search query
-Structure paper
| Title | Mapping of Ebolavirus Neutralization by Monoclonal Antibodies in the ZMapp Cocktail Using Cryo-Electron Tomography and Studies of Cellular Entry. |
|---|---|
| Journal, issue, pages | J Virol, Vol. 90, Issue 17, Page 7618-7627, Year 2016 |
| Publish date | Sep 1, 2016 |
 Authors Authors | Erin E H Tran / Elizabeth A Nelson / Pranay Bonagiri / James A Simmons / Charles J Shoemaker / Connie S Schmaljohn / Gary P Kobinger / Larry Zeitlin / Sriram Subramaniam / Judith M White /   |
| PubMed Abstract | ZMapp, a cocktail of three monoclonal antibodies (MAbs; c2G4, c4G7, and c13C6) against the ebolavirus (EBOV) glycoprotein (GP), shows promise for combatting outbreaks of EBOV, as occurred in West ...ZMapp, a cocktail of three monoclonal antibodies (MAbs; c2G4, c4G7, and c13C6) against the ebolavirus (EBOV) glycoprotein (GP), shows promise for combatting outbreaks of EBOV, as occurred in West Africa in 2014. Prior studies showed that Fabs from these MAbs bind a soluble EBOV GP ectodomain and that MAbs c2G4 and c4G7, but not c13C6, neutralize infections in cell cultures. Using cryo-electron tomography, we extended these findings by characterizing the structures of c2G4, c4G7, and c13C6 IgGs bound to native, full-length GP from the West African 2014 isolate embedded in filamentous viruslike particles (VLPs). As with the isolated ectodomain, c13C6 bound to the glycan cap, whereas c2G4 and c4G7 bound to the base region of membrane-bound GP. The tomographic data suggest that all three MAbs bind with high occupancy and that the base-binding antibodies can potentially bridge neighboring GP spikes. Functional studies indicated that c2G4 and c4G7, but not c13C6, competitively inhibit entry of VLPs bearing EBOV GP into the host cell cytoplasm, without blocking trafficking of VLPs to NPC1(+) endolysosomes, where EBOV fuses. Moreover, c2G4 and c4G7 bind to and can block entry mediated by the primed (19-kDa) form of GP without impeding binding of the C-loop of NPC1, the endolysosomal receptor for EBOV. The most likely mode of action of c2G4 and c4G7 is therefore by inhibiting conformational changes in primed, NPC1-bound GP that initiate fusion between the viral and target membranes, similar to the action of certain broadly neutralizing antibodies against influenza hemagglutinin and HIV Env. IMPORTANCE: The recent West African outbreak of ebolavirus caused the deaths of more than 11,000 individuals. Hence, there is an urgent need to be prepared with vaccines and therapeutics for similar future disasters. ZMapp, a cocktail of three MAbs directed against the ebolavirus glycoprotein, is a promising anti-ebolavirus therapeutic. Using cryo-electron tomography, we provide structural information on how each of the MAbs in this cocktail binds to the ebolavirus glycoprotein as it is displayed-embedded in the membrane and present at high density-on filamentous viruslike particles that recapitulate the surface structure and entry functions of ebolavirus. Moreover, after confirming that two of the MAbs bind to the same region in the base of the glycoprotein, we show that they competitively block the entry function of the glycoprotein and that they can do so after the glycoprotein is proteolytically primed and bound to its intracellular receptor, Niemann-Pick C1. These findings should inform future developments of ebolavirus therapeutics. |
 External links External links |  J Virol / J Virol /  PubMed:27279622 / PubMed:27279622 /  PubMed Central PubMed Central |
| Methods | EM (subtomogram averaging) |
| Resolution | 25.0 Å |
| Structure data |  EMDB-8225:  EMDB-8226: 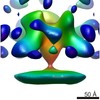 EMDB-8227:  EMDB-8228: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




 Homo sapiens (human)
Homo sapiens (human)