-検索条件
-検索結果
検索 (著者・登録者: hong & j)の結果4,232件中、1から50件目までを表示しています

EMDB-42144: 
SARS-CoV-2 Nsp15, apo-form

EMDB-42145: 
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, consensus form

EMDB-42146: 
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, state 1

EMDB-42147: 
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, state 2

PDB-8ud2: 
SARS-CoV-2 Nsp15, apo-form

PDB-8ud3: 
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, consensus form

PDB-8ud4: 
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, state 1

PDB-8ud5: 
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, state 2

EMDB-37362: 
CryoEM structure of human PI3K-alpha (P85/P110-H1047R) with QR-7909 binding at an allosteric site

EMDB-37363: 
CryoEM structure of human PI3K-alpha (P85/P110-H1047R) with QR-8557 binding at an allosteric site

PDB-8w9a: 
CryoEM structure of human PI3K-alpha (P85/P110-H1047R) with QR-7909 binding at an allosteric site

PDB-8w9b: 
CryoEM structure of human PI3K-alpha (P85/P110-H1047R) with QR-8557 binding at an allosteric site

EMDB-37154: 
Cyanophage A-1(L) neck/gp7-terminator

EMDB-37155: 
Cyanophage A-1(L) neck/gp5-neck fiber

PDB-8kef: 
Cyanophage A-1(L) neck/gp7-terminator

PDB-8keg: 
Cyanophage A-1(L) neck/gp5-neck fiber
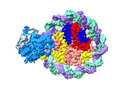
EMDB-37529: 
Structure of DDM1-nucleosome complex in the apo state
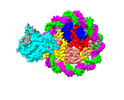
EMDB-37533: 
Structure of DDM1-nucleosome complex in ADP state
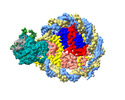
EMDB-37535: 
Structure of DDM1-nucleosome complex in ADP-BeFx state

EMDB-37537: 
Structure of DDM1-nucleosome complex in the ADP-BeFx state with DDM1 bound to SHL2 and SHL-2
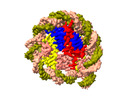
EMDB-37538: 
Structure of nucleosome core particle of Arabidopsis thaliana

PDB-8wh5: 
Structure of DDM1-nucleosome complex in the apo state

PDB-8wh8: 
Structure of DDM1-nucleosome complex in ADP state

PDB-8wh9: 
Structure of DDM1-nucleosome complex in ADP-BeFx state

PDB-8wha: 
Structure of DDM1-nucleosome complex in the ADP-BeFx state with DDM1 bound to SHL2 and SHL-2

PDB-8whb: 
Structure of nucleosome core particle of Arabidopsis thaliana
EMDB-37240: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab

EMDB-37241: 
The interface structure of Omicron RBD binding to 5817 Fab

PDB-8khc: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab

PDB-8khd: 
The interface structure of Omicron RBD binding to 5817 Fab

EMDB-38200: 
Cryo-EM structure of OSCA1.2-liposome-inside-in open state

EMDB-38503: 
Cryo-EM structure of OSCA1.2-liposome-inside-out closed state

EMDB-38611: 
Cryo-EM structure of OSCA3.1-1.1ver(Y367N-G454S-Y458I)-open/open state

EMDB-38612: 
Cryo-EM structure of OSCA3.1-1.1ver(Y367N-G454S-Y458I)-open/'desensitized' state

EMDB-38614: 
Cryo-EM structure of OSCA1.2-DOPC-1:20-contracted1 state

EMDB-38615: 
Cryo-EM structure of OSCA1.2-DOPC-1:20-contracted2 state

EMDB-38721: 
Cryo-EM structure of OSCA1.2-DOPC-1:20-expanded state

EMDB-38722: 
Cryo-EM structure of OSCA1.2-DOPC-1:50-betaCD state

EMDB-38723: 
Cryo-EM structure of OSCA3.1-2E(R611E-R619E)-closed/open state

EMDB-38724: 
Cryo-EM structure of OSCA3.1-2E(R611E-R619E)-closed/'desensitized' state

EMDB-38725: 
Cryo-EM structure of OSCA3.1-GDN state

EMDB-38726: 
Cryo-EM structure of OSCA3.1-liposome-inside-in state

EMDB-38727: 
Cryo-EM structure of OSCA1.2-V335W-DDM state

EMDB-38728: 
Cryo-EM structure of OSCA1.2-DOPC-1:50-contracted state

EMDB-38729: 
Cryo-EM structure of OSCA1.2-DOPC-1:50-expanded state

EMDB-38730: 
Cryo-EM structure of TMEM63B-Digitonin state

PDB-8xaj: 
Cryo-EM structure of OSCA1.2-liposome-inside-in open state

PDB-8xng: 
Cryo-EM structure of OSCA1.2-liposome-inside-out closed state

PDB-8xry: 
Cryo-EM structure of OSCA3.1-1.1ver(Y367N-G454S-Y458I)-open/open state

PDB-8xs0: 
Cryo-EM structure of OSCA3.1-1.1ver(Y367N-G454S-Y458I)-open/'desensitized' state
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
