-Search query
-Search result
Showing 1 - 50 of 289 items for (author: wald & j)

EMDB-17680: 
Cryo EM structure of the type 3B polymorph of alpha-synuclein at low pH.

EMDB-17693: 
Cryo EM structure of the type 3C polymorph of alpha-synuclein at low pH.

EMDB-17714: 
Cryo EM structure of the type 3D polymorph of alpha-synuclein E46K mutant at low pH.

EMDB-17723: 
Cryo EM structure of the type 1m polymorph of alpha-synuclein

EMDB-17726: 
Cryo EM structure of the type 5A polymorph of alpha-synuclein.

EMDB-50076: 
Cryo EM map of the type 2A polymorph of alpha-synuclein at pH 7.0.

EMDB-50077: 
Cryo EM map of the type 2B polymorph of alpha-synuclein at pH 7.0.

PDB-8pic: 
Cryo EM structure of the type 3B polymorph of alpha-synuclein at low pH.

PDB-8pix: 
Cryo EM structure of the type 3C polymorph of alpha-synuclein at low pH.

PDB-8pjo: 
Cryo EM structure of the type 3D polymorph of alpha-synuclein E46K mutant at low pH.

PDB-8pk2: 
Cryo EM structure of the type 1m polymorph of alpha-synuclein

PDB-8pk4: 
Cryo EM structure of the type 5A polymorph of alpha-synuclein.
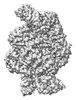
EMDB-18963: 
Structure of the SFTSV L protein in a transcription-priming state without capped RNA [TRANSCRIPTION-PRIMING (in vitro)]
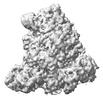
EMDB-18967: 
Structure of the SFTSV L protein in a transcription-priming state with bound capped RNA [TRANSCRIPTION-PRIMING]

EMDB-18969: 
Structure of the SFTSV L protein stalled in a transcription-specific early elongation state with bound capped RNA [TRANSCRIPTION-EARLY-ELONGATION]

PDB-8r6u: 
Structure of the SFTSV L protein in a transcription-priming state without capped RNA [TRANSCRIPTION-PRIMING (in vitro)]

PDB-8r6w: 
Structure of the SFTSV L protein in a transcription-priming state with bound capped RNA [TRANSCRIPTION-PRIMING]

PDB-8r6y: 
Structure of the SFTSV L protein stalled in a transcription-specific early elongation state with bound capped RNA [TRANSCRIPTION-EARLY-ELONGATION]

EMDB-41314: 
Structure of Gabija AB complex

EMDB-41319: 
Structure of Gabija AB complex

EMDB-41321: 
Structure of Gabija AB complex 1

PDB-8tjy: 
Structure of Gabija AB complex

PDB-8tk0: 
Structure of Gabija AB complex

PDB-8tk1: 
Structure of Gabija AB complex 1
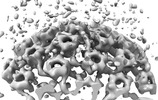
EMDB-18482: 
Herpes simplex virus 1 capsid (WT) vertices in perinuclear NEC-coated vesicles determined in situ

EMDB-18484: 
Herpes simplex virus 1 nuclear egress complex (WT) determined in situ from perinuclear vesicles

EMDB-17974: 
Pseudorabies virus cytosolic C-capsid (US3 KO) vertices determined in situ

EMDB-17975: 
Pseudorabies virus primary enveloped (perinuclear) C-capsid (US3 KO) vertices determined in situ

EMDB-17976: 
Pseudorabies nuclear C-capsids (US3 KO) vertices determined in situ

EMDB-18473: 
Subtomogram average of pseudorabies virus nuclear egress complex helical form (UL31/34) determined in situ

EMDB-18474: 
Subtomogram average of pseudorabies virus nuclear egress complex (UL31/34) determined in situ
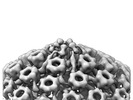

EMDB-18479: 
Pseudorabies virus cytosolic C-capsid (WT) vertices determined in situ

EMDB-18480: 
Pseudorabies virus nuclear C-capsid (WT) vertices determined in situ
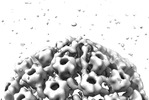

EMDB-18481: 
Herpes simplex virus 1 cytosolic C-capsid (WT) vertices determined in situ
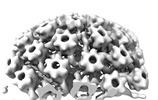
EMDB-18483: 
Herpes simplex virus 1 nuclear C-capsid (WT) vertices determined in situ

EMDB-19856: 
Focused map 1- K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide

EMDB-19857: 
Focused map 2 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide

EMDB-19858: 
Focused map 3 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide

EMDB-19859: 
Focused map 4 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide

EMDB-19860: 
Focused map 5 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide
EMDB-17798: 
CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 1

EMDB-17799: 
CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 2
EMDB-17800: 
NEDD8-CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 1

EMDB-17801: 
NEDD8-CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 2

EMDB-17802: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL1-RBX1-SKP1-FBXW7 with trapped UBE2R2-donor UB-acceptor UB-cyclin E peptide

EMDB-17803: 
Consensus map - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2-donor UB-acceptor UB-SIL1 peptide

EMDB-17822: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2-donor UB-acceptor UB-SIL1 peptide

EMDB-18767: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB~acceptor UB-BRD4

PDB-8pql: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2-donor UB-acceptor UB-SIL1 peptide
EMDB-17704: 
Subtomogram average of Vaccinia A10 trimer with open center from in vitro cores
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
