-Search query
-Search result
Showing 1 - 50 of 5,319 items for (keywords: ribosome)

EMDB entry, No image
EMDB-35903: 
The global structure of pre50S related to DbpA in state1
Method: single particle / : Yu T, Zeng F

EMDB entry, No image
EMDB-35905: 
The global structure of pre50S related to DbpA in state2
Method: single particle / : Yu T, Zeng F

EMDB entry, No image
EMDB-35908: 
The global structure of pre50S related to DbpA in state4
Method: single particle / : Yu T, Zeng F

EMDB entry, No image
EMDB-35910: 
The global structure of pre50S related to DbpA in state2
Method: single particle / : Yu T, Zeng F

EMDB entry, No image
EMDB-29897: 
Focused refinement of the 40S subunit head of the 80S Giardia lamblia ribosome at 2.94 angstroms resolution.
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB entry, No image
EMDB-29906: 
80S Giardia lamblia ribosome at 2.66 angstroms resolution with Emetine in the the V1 conformation
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-16985: 
CRYO-EM MAPS OF LEISHMANIA MAJOR 80S RIBOSOME : WILD TYPE REPLICATE 1
Method: single particle / : Rajan KS, Yonath A

EMDB-37604: 
Structural basis of translation inhibition by a valine tRNA-derived fragment
Method: single particle / : Wang YH, Zhou J

EMDB-37733: 
Structural basis of translation inhibition by a valine tRNA-derived fragment
Method: single particle / : Wang YH, Zhou J

EMDB-37734: 
Structural basis of translation inhibition by a valine tRNA-derived fragment
Method: single particle / : Wang YH, Zhou J

PDB-8wkp: 
Structural basis of translation inhibition by a valine tRNA-derived fragment
Method: single particle / : Wang YH, Zhou J

PDB-8wq2: 
Structural basis of translation inhibition by a valine tRNA-derived fragment
Method: single particle / : Wang YH, Zhou J

PDB-8wq4: 
Structural basis of translation inhibition by a valine tRNA-derived fragment
Method: single particle / : Wang YH, Zhou J

EMDB-35001: 
E. coli 70S ribosome complexed with tRNA_Ile2 bearing L34 and t6A37 in classical state
Method: single particle / : Akiyama N, Ishiguro K, Yokoyama T, Shirouzu M, Suzuki T

EMDB-35020: 
E. coli 70S ribosome complexed with H. marismortui tRNA_Ile2 bearing agm2C34 in classical state
Method: single particle / : Akiyama N, Ishiguro K, Yokoyama T, Shirouzu M, Suzuki T

EMDB-35022: 
E. coli 70S ribosome complexed with tRNA_Ile2 bearing L34 and ct6A37 in classical state
Method: single particle / : Akiyama N, Ishiguro K, Yokoyama T, Shirouzu M, Suzuki T

PDB-8hsp: 
E. coli 70S ribosome complexed with tRNA_Ile2 bearing L34 and t6A37 in classical state
Method: single particle / : Akiyama N, Ishiguro K, Yokoyama T, Shirouzu M, Suzuki T

PDB-8htz: 
E. coli 70S ribosome complexed with H. marismortui tRNA_Ile2 bearing agm2C34 in classical state
Method: single particle / : Akiyama N, Ishiguro K, Yokoyama T, Shirouzu M, Suzuki T

PDB-8hu1: 
E. coli 70S ribosome complexed with tRNA_Ile2 bearing L34 and ct6A37 in classical state
Method: single particle / : Akiyama N, Ishiguro K, Yokoyama T, Shirouzu M, Suzuki T

PDB-8rhn: 
Structure of the 55LCC ATPase complex
Method: single particle / : Foglizzo M, Degtjarik O, Zeqiraj E

EMDB-36331: 
Atomic structure of wheat ribosome reveals unique features of the plant ribosomes
Method: single particle / : Mishra RK, Sharma P, Hussain T

EMDB-36332: 
Atomic structure of wheat ribosome reveals unique features of the plant ribosomes
Method: single particle / : Mishra RK, Sharma P, Hussain T

EMDB-36333: 
Atomic structure of wheat ribosome reveals unique features of the plant ribosomes
Method: single particle / : Mishra RK, Sharma PN, Hussain T

EMDB-36334: 
Atomic structure of wheat ribosome reveals unique features of the plant ribosomes
Method: single particle / : Mishra RK, Sharma PN, Hussain T

PDB-8jiv: 
Atomic structure of wheat ribosome reveals unique features of the plant ribosomes
Method: single particle / : Mishra RK, Sharma P, Hussain T

PDB-8jiw: 
Atomic structure of wheat ribosome reveals unique features of the plant ribosomes
Method: single particle / : Mishra RK, Sharma P, Hussain T

EMDB-18534: 
Structure of SecM-stalled Escherichia coli 70S ribosome
Method: single particle / : Gersteuer F, Morici M, Wilson DN

EMDB-18590: 
Cryo-EM map of rotated SecM-stalled Escherichia coli 70S ribosome
Method: single particle / : Gersteuer F, Morici M, Wilson DN

PDB-8qoa: 
Structure of SecM-stalled Escherichia coli 70S ribosome
Method: single particle / : Gersteuer F, Morici M, Wilson DN

EMDB-18320: 
E. coli ApdP-stalled ribosomal complex
Method: single particle / : Morici M, Wilson DN

EMDB-18332: 
B. subtilis ApdA-stalled ribosomal complex
Method: single particle / : Morici M, Wilson DN

EMDB-18340: 
ApdP-SRC with P-tRNA only
Method: single particle / : Morici M, Wilson DN

EMDB-18341: 
ApdA-SRC with P-tRNA only
Method: single particle / : Morici M, Wilson DN

EMDB-17329: 
48S late-stage initiation complex with m6A mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

EMDB-17330: 
48S late-stage initiation complex with non methylated mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

PDB-8p03: 
48S late-stage initiation complex with m6A mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

PDB-8p09: 
48S late-stage initiation complex with non methylated mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

EMDB-43851: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP63 Class E2
Method: single particle / : Dong X, Sheng K, Williamson JR

EMDB-43853: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP63 Class D
Method: single particle / : Dong X, Sheng K, Williamson JR

EMDB-43854: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP63 Class B
Method: single particle / : Dong X, Sheng K, Williamson JR
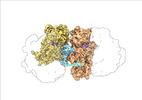
EMDB-19054: 
Escherichia coli paused disome complex (Non-rotated disome interface class 1)
Method: single particle / : Fluegel T, Schacherl M

PDB-8rcl: 
Escherichia coli paused disome complex (Non-rotated disome interface class 1)
Method: single particle / : Fluegel T, Schacherl M

EMDB-43846: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP45 Class C2
Method: single particle / : Dong X, Sheng K, Williamson JR

EMDB-43847: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP45 Class E2
Method: single particle / : Dong X, Sheng K, Williamson JR

EMDB-43848: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP45 Class G
Method: single particle / : Dong X, Sheng K, Williamson JR

EMDB-43849: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP45 Class E1
Method: single particle / : Dong X, Sheng K, Williamson JR

EMDB-43856: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP63(+PS) Class E
Method: single particle / : Dong X, Sheng K, Williamson JR

EMDB-43857: 
Structure of Circularly Permuted 50S Ribosomal Subunit Assembly Intermediate - CP63(+PS) Class C2
Method: single particle / : Dong X, Sheng K, Williamson JR
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
