-Search query
-Search result
Showing all 33 items for (author: de & la & cruz & mj)

EMDB-25132: 
Androgen receptor bound to DNA - Entrenched state
Method: single particle / : Wasmuth EV, Vanden Broeck A, Klinge S, Sawyers CL

EMDB-25133: 
Androgen receptor bound to DNA - Splayed state
Method: single particle / : Wasmuth EV, Vanden Broeck A, Klinge S, Sawyers CL

EMDB-25134: 
Androgen receptor bound to DNA - Divorced state
Method: single particle / : Wasmuth EV, Vanden Broeck A, Klinge S, Sawyers CL

EMDB-24292: 
Cryo-EM structure of DNMT5 in apo state
Method: single particle / : Wang J, Patel DJ

EMDB-24294: 
Cryo-EM structure of DNMT5 binary complex with hemimethylated DNA
Method: single particle / : Wang J, Patel DJ

EMDB-24295: 
cryo-EM structure of DNMT5 quaternary complex with hemimethylated DNA, AMP-PNP and SAH
Method: single particle / : Wang J, Patel DJ

EMDB-25577: 
Cryo-EM structure of DNMT5 pseudo-ternary complex solved by incubation with hemimethylated DNA and SAM
Method: single particle / : Wang J, Patel DJ

EMDB-23282: 
polynucleotide phosphorylase
Method: single particle / : Goldgur Y, Shuman S, De La Cruz MJ, Ghosh S, Unciuleac MC

EMDB-22170: 
BG505 SOSIPv5.2 in complex with PGT122 and two RM20E1 Fabs
Method: single particle / : Cottrell CA, Ward AB

EMDB-22171: 
BG505 SOSIPv5.2 in complex with PGT122 and three RM20E1 Fabs
Method: single particle / : Cottrell CA, Ward AB

EMDB-22172: 
C3 symmetric BG505 SOSIPv5.2 in complex with PGT122 and three RM20E1 Fabs
Method: single particle / : Cottrell CA, Ward AB

EMDB-22173: 
BG505 SOSIPv5.2 in complex with PGT122 and one RM20E1 Fab
Method: single particle / : Cottrell CA, Ward AB

EMDB-22178: 
BG505 SOSIPv5.2 in complex with PGT122 and no RM20E1 Fabs
Method: single particle / : Cottrell CA, Ward AB

EMDB-22179: 
C3 symmetric BG505 SOSIPv5.2 in complex with PGT122 and no RM20E1 Fabs
Method: single particle / : Cottrell CA, Ward AB

EMDB-23244: 
Structure of human SHLD2-SHLD3-REV7-TRIP13(E253Q) complex
Method: single particle / : Xie W, Patel DJ

EMDB-21126: 
Cryo-EM structure of Cascade-TniQ binary complex
Method: single particle / : Jia N, Patel DJ

EMDB-21146: 
Cryo-EM structure of Cascade-TniQ-dsDNA ternary complex
Method: single particle / : Jia N, Patel DJ

EMDB-7321: 
Integrative Structure and Functional Anatomy of a Nuclear Pore Complex
Method: subtomogram averaging / : Kim SJ, Fernandez-Martinez J, Nudelman I, Shi Y, Zhang W, Ludtke SJ, Akey CW, Chait BT, Sali A, Rout MP

EMDB-8216: 
MicroED structure of tau VQIVYK peptide at 1.1 A resolution
Method: electron crystallography / : de la Cruz MJ, Hattne J

EMDB-8217: 
MicroED structure of lysozyme at 1.8 A resolution
Method: electron crystallography / : de la Cruz MJ, Hattne J, Shi D, Seidler P, Rodriguez J, Reyes FE, Sawaya MR, Cascio D, Eisenberg D, Gonen T

EMDB-8218: 
MicroED structure of xylanase at 2.3 A resolution
Method: electron crystallography / : de la Cruz MJ, Hattne J

EMDB-8219: 
MicroED structure of thaumatin at 2.5 A resolution
Method: electron crystallography / : de la Cruz MJ, Hattne J, Shi D, Seidler P, Rodriguez J, Reyes FE, Sawaya MR, Cascio D, Eisenberg D, Gonen T

EMDB-8220: 
MicroED structure of trypsin at 1.7 A resolution
Method: electron crystallography / : de la Cruz MJ, Hattne J, Shi D, Seidler P, Rodriguez J, Reyes FE, Sawaya MR, Cascio D, Eisenberg D, Gonen T

EMDB-8221: 
MicroED structure of proteinase K at 1.6 A resolution
Method: electron crystallography / : de la Cruz MJ, Hattne J, Shi D, Seidler P, Rodriguez J, Reyes FE, Sawaya MR, Cascio D, Eisenberg D, Gonen T

EMDB-8222: 
MicroED structure of thermolysin at 2.5 A resolution
Method: electron crystallography / : de la Cruz MJ, Hattne J

EMDB-8472: 
MicroED structure of a complex between monomeric TGF-b and its receptor, TbRII, at 2.9 A resolution
Method: electron crystallography / : Weiss SC, de la Cruz MJ

EMDB-8077: 
MicroED structure of proteinase K at 1.75 A resolution
Method: electron crystallography / : Hattne J, Shi D

EMDB-6428: 
Cryo-EM structure of MAVS CARD C1 filament
Method: helical / : Xu H, He X, Zheng H, Huang LJ, Hou F, Yu Z, de la Cruz MJ, Borkowski B, Zhang X, Chen ZJ, Jiang QX

EMDB-5890: 
Cryo-EM structure of MAVS CARD filament
Method: helical / : Xu H, He X, Zheng H, Huang LJ, Hou F, Yu Z, de la Cruz MJ, Borkowski B, Zhang X, Chen ZJ, Jiang QX

EMDB-5891: 
Cryo-EM structure of MAVSdeltaProTM filament
Method: helical / : Xu H, He X, Zheng H, Huang LJ, Hou F, Yu Z, de la Cruz MJ, Borkowski B, Zhang X, Chen ZJ, Jiang QX

EMDB-5272: 
Molecular Structure of Unliganded Native SIVmneE11S gp120 trimer: Spike region
Method: subtomogram averaging / : White TA, Bartesaghi A, Borgnia M, Subramaniam S
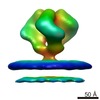
EMDB-5273: 
Molecular Structure of Unliganded Native SIVmac239 gp120 trimer: Spike region
Method: subtomogram averaging / : White TA, Bartesaghi A, Borgnia M, Subramaniam S

EMDB-5274: 
Molecular Structure of Unliganded Native CP-MAC gp120 trimer: Spike region
Method: subtomogram averaging / : White TA, Bartesaghi A, Borgnia M, Subramaniam S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
