-Search query
-Search result
Showing 1 - 50 of 4,020 items for (author: zhao & c)
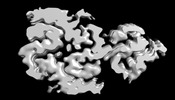
EMDB-37197: 
The cryo-EM structure of type3 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37200: 
The cryo-EM structure of AV-45 bound type3 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37199: 
The cryo-EM structure of type1 amyloid beta 42 fibril in AD3.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37195: 
The cryo-EM structure of AV-45 bound type1 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-60099: 
SARS-CoV-2 spike trimer (6P) in complex with two R1-26 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60100: 
SARS-CoV-2 spike trimer (6P) in complex with three R1-26 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60101: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, head-to-head aggregate
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60102: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, focused refinement of RBD-Fab region
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60103: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60104: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60105: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C1 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60106: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C3 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60107: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 and two R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60108: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60109: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs (one RBD rotated)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60110: 
SARS-CoV-2 S1 in complex with H18 and R1-32 Fab
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60111: 
Dimer of SARS-CoV-2 S1 in complex with H18 and R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-37198: 
The cryo-EM structure of type1 amyloid beta 42 fibril in AD2 patient.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-38389: 
Cryo-EM structure of sheep VMAT2 dimer in an atypical fold
Method: single particle / : Lyu Y, Fu C, Ma H, Sun Z, Su Z, Zhou X
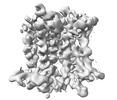
EMDB-38390: 
Cryo-EM structure of a frog VMAT2 in an apo conformation
Method: single particle / : Lyu Y, Fu C, Ma H, Sun Z, Su Z, Zhou X

PDB-8xit: 
Cryo-EM structure of sheep VMAT2 dimer in an atypical fold
Method: single particle / : Lyu Y, Fu C, Ma H, Sun Z, Su Z, Zhou X

PDB-8xiu: 
Cryo-EM structure of a frog VMAT2 in an apo conformation
Method: single particle / : Lyu Y, Fu C, Ma H, Sun Z, Su Z, Zhou X

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38846: 
Block 1 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38848: 
Block 2 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-43814: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in post-cleavage stage
Method: single particle / : Wang B, Goswami HN, Li H

EMDB-43815: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in pre-cleavage stage
Method: single particle / : Wang B, Goswami HN, Li H

PDB-9ash: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in post-cleavage stage
Method: single particle / : Wang B, Goswami HN, Li H

PDB-9asi: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in pre-cleavage stage
Method: single particle / : Wang B, Goswami HN, Li H

EMDB-41854: 
Structure of Human Mitochondrial Chaperonin V72I Mutant
Method: single particle / : Chen L, Wang J

PDB-8u39: 
Structure of Human Mitochondrial Chaperonin V72I mutant
Method: single particle / : Chen L, Wang J
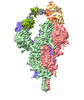
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
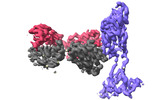
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
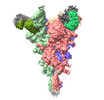
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
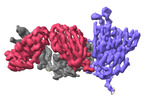

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
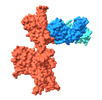
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
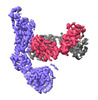
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
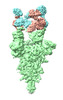
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
