-Search query
-Search result
Showing 1 - 50 of 4,087 items for (author: yang & k)

EMDB entry, No image
EMDB-43931: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

EMDB entry, No image
EMDB-43932: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axa: 
CryoEM structure of activated CRAF/MEK/14-3-3 complex with NST-628
Method: single particle / : Quade B, Cohen SE, Huang X

PDB-9axc: 
Activated CRAF/MEK heterotetramer from focused refinement of CRAF/MEK/14-3-3 complex
Method: single particle / : Quade B, Cohen SE, Huang X

EMDB entry, No image
EMDB-43705: 
HIV-1 wild-type intasome core
Method: single particle / : Li M, Craigie R

EMDB entry, No image
EMDB-43756: 
HIV-1 P5-IN intasome core
Method: single particle / : Li M, Craigie R

EMDB entry, No image
EMDB-43761: 
HIV-1 intasome core assembled with wild-type integrase, 1F
Method: single particle / : Li M, Craigie R

PDB-8w34: 
HIV-1 intasome core assembled with wild-type integrase, 1F
Method: single particle / : Li M, Craigie R

EMDB entry, No image
EMDB-37154: 
Cyanophage A-1(L) neck/gp7-terminator
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB entry, No image
EMDB-37155: 
Cyanophage A-1(L) neck/gp5-neck fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB entry, No image
EMDB-37240: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab
Method: single particle / : Cao L, Wang X

EMDB entry, No image
EMDB-37241: 
The interface structure of Omicron RBD binding to 5817 Fab
Method: single particle / : Cao L, Wang X

PDB-8khc: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab
Method: single particle / : Cao L, Wang X

PDB-8khd: 
The interface structure of Omicron RBD binding to 5817 Fab
Method: single particle / : Cao L, Wang X

EMDB-37389: 
cryo-EM structure of native mastigonemes isolated from Chlamydomonas reinhardtii at 3.0 angstrom resolution
Method: single particle / : Huang J, Tao H, Chen J, Pan J, Yan C, Yan N

PDB-8wa2: 
cryo-EM structure of native mastigonemes isolated from Chlamydomonas reinhardtii at 3.0 angstrom resolution
Method: single particle / : Huang J, Tao H, Chen J, Pan J, Yan C, Yan N

EMDB-37150: 
The CBD domain of cyanophage A-1(L) short tail fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37151: 
Cyanophage A-1(L) baseplate-initiators
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37152: 
Cyanophage A-1(L) tail fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-37153: 
Cyanophage A-1(L) sheath-tube
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-41590: 
Cyanophage A-1(L) portal
Method: single particle / : Yu RC, Li Q, Zhou CZ

PDB-8ke9: 
The CBD domain of cyanophage A-1(L) short tail fiber
Method: single particle / : Yu RC, Li Q, Zhou CZ

EMDB-43008: 
Fab fragment of human mAb #58 in complex with computationally optimized broadly reactive H1 influenza hemagglutinin X6
Method: single particle / : Nagashima KA, Mousa JJ

EMDB-35990: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-translocation state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

EMDB-35991: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the resting state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

EMDB-35992: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-catalytic intermediate state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

EMDB-35993: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the catalytic intermediate state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

PDB-8j5q: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-translocation state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

PDB-8j5r: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the resting state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

PDB-8j5s: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-catalytic intermediate state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

PDB-8j5t: 
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the catalytic intermediate state
Method: single particle / : Yang X, Hu T, Zhang B, Rao Z

EMDB-36212: 
PhK holoenzyme in inactive state, muscle isoform
Method: single particle / : Yang XK, Xiao JY

EMDB-36213: 
PhK holoenzyme in active state, muscle isoform
Method: single particle / : Yang XK, Xiao JY

EMDB-36214: 
local map of hPhK alpha-beta-gamma-delta subcomplex in inactive state
Method: single particle / : Yang XK, Xiao JY

EMDB-36215: 
local map of hPhK gamma-delta subcomplex in inactive state
Method: single particle / : Yang XK, Xiao JY

EMDB-36216: 
local map of hPhK alpha-gamma subcomplex in active state
Method: single particle / : Yang XK, Xiao JY

PDB-8jfk: 
PhK holoenzyme in inactive state, muscle isoform
Method: single particle / : Yang XK, Xiao JY

PDB-8jfl: 
PhK holoenzyme in active state, muscle isoform
Method: single particle / : Yang XK, Xiao JY

PDB-8xya: 
hPhK alpha-beta-gamma-delta subcomplex in inactive state
Method: single particle / : Yang XK, Xiao JY
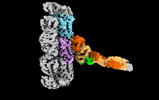
EMDB-18304: 
Outer kinetochore Ndc80-Dam1 alpha/beta-tubulin complex
Method: single particle / : Muir KW, Barford D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search













 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
