-Search query
-Search result
Showing 1 - 50 of 2,377 items for (author: yan & qin)

EMDB-31367: 
Structure of mumps virus nucleoprotein without C-arm
Method: helical / : Shen Q, Shan H, Zhang N, Qin Y

PDB-7exa: 
Structure of mumps virus nucleoprotein without C-arm
Method: helical / : Shen Q, Shan H, Zhang N, Qin Y

EMDB-38850: 
Cryo-EM structure of human dopamine transporter in apo state
Method: single particle / : Zhao Y, Li Y

EMDB-38851: 
Cryo-EM structure of human dopamine transporter in complex with dopamine
Method: single particle / : Zhao Y, Li Y

EMDB-38852: 
Cryo-EM structure of human dopamine transporter in complex with benztropine
Method: single particle / : Zhao Y, Li Y

EMDB-38853: 
structure of a proteinACryo-EM structure of human dopamine transporter in complex with GBR12909
Method: single particle / : Zhao Y, Li Y

EMDB-38854: 
Cryo-EM structure of human dopamine transporter in complex with methylphenidate
Method: single particle / : Zhao Y, Li Y

PDB-8y2c: 
Cryo-EM structure of human dopamine transporter in apo state
Method: single particle / : Zhao Y, Li Y

PDB-8y2d: 
Cryo-EM structure of human dopamine transporter in complex with dopamine
Method: single particle / : Zhao Y, Li Y

PDB-8y2e: 
Cryo-EM structure of human dopamine transporter in complex with benztropine
Method: single particle / : Zhao Y, Li Y

PDB-8y2f: 
Cryo-EM structure of human dopamine transporter in complex with GBR12909
Method: single particle / : Zhao Y, Li Y

PDB-8y2g: 
Cryo-EM structure of human dopamine transporter in complex with methylphenidate
Method: single particle / : Zhao Y, Li Y

EMDB-38300: 
Cryo-EM structure of partial dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39863: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39943: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body2
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39947: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body1
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39949: 
Cryo-EM structure of WDR11-dm-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

PDB-8xfb: 
Cryo-EM structure of partial dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

PDB-8z9m: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-37842: 
Cryo-EM structure of noradrenaline transporter in apo state
Method: single particle / : Zhao Y, Hu T, Yu Z
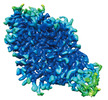
EMDB-37843: 
Cryo-EM structure of noradrenaline transporter in complex with noradrenaline
Method: single particle / : Zhao Y, Hu T, Yu Z
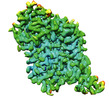
EMDB-37844: 
Cryo-EM structure of noradrenaline transporter in complex with a x-MrlA analogue
Method: single particle / : Zhao Y, Hu T, Yu Z
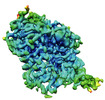
EMDB-37845: 
Cryo-EM structure of noradrenaline transporter in complex with bupropion
Method: single particle / : Zhao Y, Hu T, Yu Z
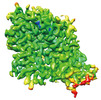
EMDB-37846: 
Cryo-EM structure of noradrenaline transporter in complex with ziprasidone
Method: single particle / : Zhao Y, Hu T, Yu Z

PDB-8wtu: 
Cryo-EM structure of noradrenaline transporter in apo state
Method: single particle / : Zhao Y, Hu T, Yu Z

PDB-8wtv: 
Cryo-EM structure of noradrenaline transporter in complex with noradrenaline
Method: single particle / : Zhao Y, Hu T, Yu Z

PDB-8wtw: 
Cryo-EM structure of noradrenaline transporter in complex with a x-MrlA analogue
Method: single particle / : Zhao Y, Hu T, Yu Z

PDB-8wtx: 
Cryo-EM structure of noradrenaline transporter in complex with bupropion
Method: single particle / : Zhao Y, Hu T, Yu Z

PDB-8wty: 
Cryo-EM structure of noradrenaline transporter in complex with ziprasidone
Method: single particle / : Zhao Y, Hu T, Yu Z

EMDB-39623: 
Cryo-EM structure of the retatrutide-bound human GCGR-Gs complex
Method: single particle / : Li WZ, Zhou QT, Cong ZT, Yuan QN, Li WX, Zhao FH, Xu HE, Zhao LH, Yang DH, Wang MW

PDB-8yw5: 
Cryo-EM structure of the retatrutide-bound human GCGR-Gs complex
Method: single particle / : Li WZ, Zhou QT, Cong ZT, Yuan QN, Li WX, Zhao FH, Xu HE, Zhao LH, Yang DH, Wang MW

EMDB-39034: 
Human AE3 with NaHCO3- and DIDS
Method: single particle / : Jian L, Zhang Q, Yao D, Wang Q, Xia Y, Qin A, Cao Y

EMDB-39035: 
Human AE3 with NaHCO3-
Method: single particle / : Jian L, Zhang Q, Yao D, Cao Y

EMDB-39050: 
The structure of hAE3
Method: single particle / : Jian L, Zhang Q, Yao D, Wang Q, Xia Y, Qin A, Cao Y

EMDB-60225: 
hAE3NTD2TMD with PT5,CLR, and Y01
Method: single particle / : Jian L, Zhang Q, Yao D, Wang Q, Xia Y, Qin A, Cao Y

PDB-8y85: 
Human AE3 with NaHCO3- and DIDS
Method: single particle / : Jian L, Zhang Q, Yao D, Wang Q, Xia Y, Qin A, Cao Y

PDB-8y8k: 
The structure of hAE3
Method: single particle / : Jian L, Zhang Q, Yao D, Wang Q, Xia Y, Qin A, Cao Y

PDB-8zle: 
hAE3NTD2TMD with PT5,CLR, and Y01
Method: single particle / : Jian L, Zhang Q, Yao D, Wang Q, Xia Y, Qin A, Cao Y

EMDB-60607: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-60608: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Ggust
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-60626: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH
EMDB-60627: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9iiw: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9iix: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Ggust
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9ij9: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9ija: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-32979: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)
Method: single particle / : Zheng Q, Zhu R, Sun H, Cheng T, Li S, Xia N

PDB-7x35: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)
Method: single particle / : Zheng Q, Zhu R, Sun H, Cheng T, Li S, Xia N

EMDB-37513: 
Cryo-EM structure of the red-shifted Fittonia albivenis PSI-LHCI
Method: single particle / : Huang GQ, Li XX, Sui SF, Qin XC
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
