-Search query
-Search result
Showing 1 - 50 of 92 items for (author: wu & cc)

EMDB-41374: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS
EMDB-41382: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS
EMDB-41399: 
Antibody N3-1 bound to SARS-CoV-2 spike
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tm1: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tma: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS

EMDB-34490: 
The cryo-EM structure of nuclear transport receptor Kap114p complex with yeast TATA-box binding protein
Method: single particle / : Hsia KC, Liao CC, Wang CH, Wu YM

EMDB-34935: 
Cryo-EM structure of a SIN3/HDAC complex from budding yeast
Method: single particle / : Guo Z, Zhan X, Wang C
EMDB-34936: 
Cryo-EM map of the Rpd3L complex from the Rxt1-Flag dataset
Method: single particle / : Guo Z, Zhan X, Wang C

EMDB-34937: 
Cryo-EM map of the Rpd3L complex in Sin3 part
Method: single particle / : Guo Z, Zhan X, Wang C

EMDB-34938: 
Cryo-EM map of the Rpd3L complex in Dep1 region
Method: single particle / : Guo Z, Zhan X, Wang C
EMDB-34939: 
Cryo-EM map of the Rpd3L complex in Ume1 part
Method: single particle / : Guo Z, Zhan X, Wang C

EMDB-27704: 
Cryo-EM structure of insulin receptor (IR) bound with S597 peptide
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

EMDB-27705: 
Cryo-EM structure of insulin receptor (IR) bound with S597 component 2
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

PDB-8dtl: 
Cryo-EM structure of insulin receptor (IR) bound with S597 peptide
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

PDB-8dtm: 
Cryo-EM structure of insulin receptor (IR) bound with S597 component 2
Method: single particle / : Park J, Li J, Mayer JP, Ball KA, Wu JY, Hall C, Accili D, Stowell MHB, Bai XC, Choi E

EMDB-32243: 
Cryo-EM structure of a dimeric GPCR-Gi complex with small molecule
Method: single particle / : Yue Y, Liu LE, Wu LJ, Xu F, Hanson M

EMDB-32244: 
Cryo-EM structure of a monomeric GPCR-Gi complex with small molecule
Method: single particle / : Xu F, Yue Y, Liu LE, Wu LJ, Hanson M

EMDB-32245: 
Cryo-EM structure of a dimeric GPCR-Gi complex with peptide
Method: single particle / : Xu F, Yue Y, Wu LJ, Liu LE, Hanson M

EMDB-32246: 
Cryo-EM structure of a monomeric GPCR-Gi complex with peptide
Method: single particle / : Xu F, Yue Y, Liu LE, Wu LJ, Hanson M

EMDB-32247: 
Cryo-EM structure of a GPCR-Gi complex with peptide
Method: single particle / : Xu F, Yue Y, Liu LE, Wu LJ, Hanson M

PDB-7w0l: 
Cryo-EM structure of a dimeric GPCR-Gi complex with small molecule
Method: single particle / : Yue Y, Liu LE, Wu LJ, Xu F, Hanson M

PDB-7w0m: 
Cryo-EM structure of a monomeric GPCR-Gi complex with small molecule
Method: single particle / : Xu F, Yue Y, Liu LE, Wu LJ, Hanson M

PDB-7w0n: 
Cryo-EM structure of a dimeric GPCR-Gi complex with peptide
Method: single particle / : Xu F, Yue Y, Wu LJ, Liu LE, Hanson M

PDB-7w0o: 
Cryo-EM structure of a monomeric GPCR-Gi complex with peptide
Method: single particle / : Xu F, Yue Y, Liu LE, Wu LJ, Hanson M

PDB-7w0p: 
Cryo-EM structure of a GPCR-Gi complex with peptide
Method: single particle / : Xu F, Yue Y, Liu LE, Wu LJ, Hanson M

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
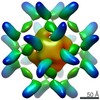
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-24362: 
Cryo-EM structure of M4008_N1 Fab in complex with BG505 DS-SOSIP.664 Env trimer
Method: single particle / : Chan KW, Kong XP

EMDB-31470: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-25 (Focused refinement of S-RBD and chAb-25 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD

EMDB-31471: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-45 (Focused refinement of S-RBD and chAb-45 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD

EMDB-23494: 
Cryo-EM of the SLFN12-PDE3A complex: PDE3A body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

EMDB-23495: 
Cryo-EM of the SLFN12-PDE3A complex: Consensus subset model
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

EMDB-23496: 
Cryo-EM of the SLFN12-PDE3A complex: SLFN12 body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

PDB-7lrc: 
Cryo-EM of the SLFN12-PDE3A complex: PDE3A body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

PDB-7lrd: 
Cryo-EM of the SLFN12-PDE3A complex: Consensus subset model
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

PDB-7lre: 
Cryo-EM of the SLFN12-PDE3A complex: SLFN12 body refinement
Method: single particle / : Fuller JR, Garvie CW, Lemke CT

EMDB-22490: 
Structure of human TRPA1 in complex with antagonist compound 21
Method: single particle / : Rohou A, Rouge L, Chen H

PDB-7jup: 
Structure of human TRPA1 in complex with antagonist compound 21
Method: single particle / : Rohou A, Rouge L

EMDB-12059: 
Yeast 80S ribosome bound to eEF3 and A/A- and P/P-tRNAs (PRE-1)
Method: single particle / : Ranjan N, Pochopien AA, Wu CC, Beckert B, Blanchet S, Green R, Rodnina MV, Wilson DN

EMDB-12062: 
Yeast 80S ribosome bound to eEF3 and P/P- and E/E-tRNAs (POST-2)
Method: single particle / : Ranjan N, Pochopien AA, Wu CC, Beckert B, Blanchet S, Green R, Rodnina MV, Wilson DN

EMDB-12064: 
Yeast 80S ribosome bound to eEF3 and P/P-tRNA (POST-3)
Method: single particle / : Ranjan N, Pochopien AA, Wu CC, Beckert B, Blanchet S, Green R, Rodnina MV, Wilson DN

EMDB-12065: 
Yeast 80S ribosome with bound A/P- and P/E-tRNAs (PRE-4)
Method: single particle / : Ranjan N, Pochopien AA, Wu CC, Beckert B, Blanchet S, Green R, Rodnina MV, Wilson DN

EMDB-12074: 
Yeast 80S ribosome bound to eEF2 and ap/P- and pe/E-tRNAs (POST-1).
Method: single particle / : Ranjan N, Pochopien AA, Wu CC, Beckert B, Blanchet S, Green R, Rodnina MV, Wilson DN

EMDB-12075: 
Yeast 80S ribosome with bound A/A- and P/E-tRNAs (PRE-3).
Method: single particle / : Ranjan N, Pochopien AA, Wu CC, Beckert B, Blanchet S, Green R, Rodnina MV, Wilson DN

EMDB-12081: 
Yeast 80S ribosome bound to eEF3 and A/A- and P/P-tRNAs
Method: single particle / : Ranjan N, Pochopien AA, Wu CC, Beckert B, Blanchet S, Green R, Rodnina MV, Wilson DN

EMDB-30392: 
Cryo-EM structure of Fenoldopam bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao W

EMDB-30393: 
Cryo-EM structure of A77636 bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao Z

EMDB-30394: 
Cryo-EM structure of PW0464 bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao Z
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
