-Search query
-Search result
Showing 1 - 50 of 64 items for (author: walter & ts)

EMDB-41940: 
Human RADX tetramer bound to ssDNA
Method: single particle / : Balakrishnan S, Chazin WJ

PDB-8u61: 
Human RADX tetramer bound to ssDNA
Method: single particle / : Balakrishnan S, Chazin WJ

EMDB-41617: 
CryoEM structure of PI3Kalpha
Method: single particle / : Valverde R, Shi H, Holliday M, Sun M

PDB-8tu6: 
CryoEM structure of PI3Kalpha
Method: single particle / : Valverde R, Shi H, Holliday M

EMDB-41939: 
human RADX trimer bound to ssDNA
Method: single particle / : Balakrishnan S, Chazin WJ

EMDB-42156: 
RADX trimer locally refined map 1
Method: single particle / : Balakrishnan S, Chazin WJ

EMDB-42157: 
RADX trimer locally refined map 2
Method: single particle / : Balakrishnan S, Chazin WJ

EMDB-42162: 
RADX trimer consensus map
Method: single particle / : Balakrishnan S, Chazin WJ

PDB-8u5y: 
human RADX trimer bound to ssDNA
Method: single particle / : Balakrishnan S, Chazin WJ

EMDB-26160: 
Structure of human serotonin transporter bound to small molecule '8090 in lipid nanodisc and NaCl
Method: single particle / : Singh I, Seth A

PDB-7txt: 
Structure of human serotonin transporter bound to small molecule '8090 in lipid nanodisc and NaCl
Method: single particle / : Singh I, Seth A, Billesboelle CB, Braz J, Rodriguiz RM, Roy K, Bekele B, Craik V, Huang XP, Boytsov D, Lak P, O'Donnell H, Sandtner W, Roth BL, Basbaum AI, Wetsel WC, Manglik A, Shoichet BK, Rudnick G

EMDB-25570: 
Cryo-EM structure of Rifamycin bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS

EMDB-25571: 
Cryo-EM structure of 27a bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS

PDB-7szj: 
Cryo-EM structure of Rifamycin bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS

PDB-7szk: 
Cryo-EM structure of 27a bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS
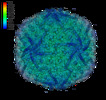
EMDB-13109: 
Bacteriophage PRD1 in complex with chlorpromazine (CPZ)
Method: single particle / : Duyvesteyn HME, Santos-Perez I, Peccati F, Martinez-Castillo A, Jimenez-Oses G, Oksanen HM, Stuart DI, Abrescia NGA

EMDB-24334: 
cryo-EM of human Gastric inhibitory polypeptide receptor GIPR bound to GIP
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

EMDB-24401: 
cryo-EM structure of human Gastric inhibitory polypeptide receptor GIPR bound to tirzepatide
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

EMDB-24445: 
cryo-EM of human Glucagon-like peptide 1 receptor GLP-1R in apo form
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

EMDB-24453: 
cryo-EM of human Glucagon-like peptide 1 receptor GLP-1R bound to tirzepatide
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

PDB-7ra3: 
cryo-EM of human Gastric inhibitory polypeptide receptor GIPR bound to GIP
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

PDB-7rbt: 
cryo-EM structure of human Gastric inhibitory polypeptide receptor GIPR bound to tirzepatide
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

PDB-7rg9: 
cryo-EM of human Glucagon-like peptide 1 receptor GLP-1R in apo form
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

PDB-7rgp: 
cryo-EM of human Glucagon-like peptide 1 receptor GLP-1R bound to tirzepatide
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

PDB-7l7g: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B from Homo sapiens (updated model of PDB ID: 6CAJ)
Method: single particle / : Tsai JC, Miller-Vedam LE, Anand A, Jaishankar P, Nguyen HC, Wang L, Renslo AR, Frost A, Walter P

EMDB-12274: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12275: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-88 Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12276: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-150 Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12277: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12278: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-316 Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12279: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-384 Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12280: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-253H55L Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12281: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-253H55L Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12282: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-253H165L Fab
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12283: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-159
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-12284: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-159
Method: single particle / : Duyvesteyn HME, Zhao Y, Ren J, Stuart D

EMDB-22283: 
cryo-EM of human GLP-1R bound to non-peptide agonist LY3502970
Method: single particle / : Sun B, Kobilka BK, Sloop KW, Feng D, Kobilka TS

EMDB-21147: 
Cryo-EM structure of the Glucagon-like peptide-1 receptor in complex with G protein, GLP-1 peptide and a positive allosteric modulator
Method: single particle / : Sun B, Feng D, Bueno A, Kobilka B, Sloop K

PDB-6vcb: 
Cryo-EM structure of the Glucagon-like peptide-1 receptor in complex with G protein, GLP-1 peptide and a positive allosteric modulator
Method: single particle / : Sun B, Feng D, Bueno A, Kobilka B, Sloop K

EMDB-0649: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B bound to translation initiation factor 2 from Homo sapiens
Method: single particle / : Nguyen H, Kenner L

EMDB-0651: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B bound to eukaryotic translation initiation factor 2 from Homo sapiens
Method: single particle / : Nguyen HC, Kenner LR

EMDB-0664: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B bound to eukaryotic translation initiation factor 2 from Homo sapiens
Method: single particle / : Nguyen HC, Kenner LR, Frost AS

PDB-6o81: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B bound to translation initiation factor 2 from Homo sapiens
Method: single particle / : Nguyen H, Kenner L, Frost A

PDB-6o85: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B bound to eukaryotic translation initiation factor 2 from Homo sapiens
Method: single particle / : Nguyen HC, Kenner LR, Frost AS

PDB-6o9z: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B bound to eukaryotic translation initiation factor 2 from Homo sapiens
Method: single particle / : Nguyen HC, Kenner LR, Frost AS

EMDB-4633: 
Cryo-EM structure of SH1 full particle.
Method: single particle / : De Colibus L, Roine E, Walter TS, Ilca SL, Wang X, Wang N, Roseman AM, Bamford D, Huiskonen JT, Stuart DI

EMDB-4634: 
Localized reconstruction of archaeal virus SH1 vertex calculated without symmetry
Method: single particle / : De Colibus L, Ilca SL, Stuart DIS, Huiskonen JT

EMDB-4656: 
Localized reconstruction of archaeal virus SH1 spike calculated with C2 symmetry
Method: single particle / : De Colibus L, Ilca SL, Stuart DIS, Huiskonen JT

EMDB-9779: 
Reconstruction of HRPV6 VP5 spike
Method: subtomogram averaging / : Li S, Huiskonen JT

EMDB-7442: 
Electron cryo-microscopy of the eukaryotic translation initiation factor 2B from Homo sapiens
Method: single particle / : Tsai JC, Miller-Vedam LE, Anand AA, Jaishankar P, Nguyen HC, Renslo AR, Frost A, Walter P
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
