-Search query
-Search result
Showing 1 - 50 of 1,335 items for (author: tong & w)

EMDB-42291: 
Structure of the human INTS9-INTS11-BRAT1 complex
Method: single particle / : Lin M, Tong L

EMDB-42292: 
Structure of the Drosophila IntS11-CG7044(dBRAT1) complex
Method: single particle / : Lin M, Tong L

PDB-8uib: 
Structure of the human INTS9-INTS11-BRAT1 complex
Method: single particle / : Lin M, Tong L

PDB-8uic: 
Structure of the Drosophila IntS11-CG7044(dBRAT1) complex
Method: single particle / : Lin M, Tong L

EMDB-39623: 
Cryo-EM structure of the retatrutide-bound human GCGR-Gs complex
Method: single particle / : Li WZ, Zhou QT, Cong ZT, Yuan QN, Li WX, Zhao FH, Xu HE, Zhao LH, Yang DH, Wang MW

PDB-8yw5: 
Cryo-EM structure of the retatrutide-bound human GCGR-Gs complex
Method: single particle / : Li WZ, Zhou QT, Cong ZT, Yuan QN, Li WX, Zhao FH, Xu HE, Zhao LH, Yang DH, Wang MW
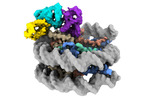
EMDB-38099: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by intein-based E2-Ub-NCP conjugation strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
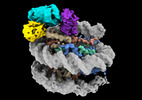
EMDB-38100: 
Cryo-EM structures of RNF168/UbcH5c-Ub/nucleosomes complex determined by activity-based chemical trapping strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
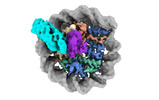
EMDB-38101: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by activity-based chemical trapping strategy (adjacent H2AK13/15 dual-monoubiquitination)
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
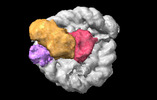
EMDB-38102: 
Cryo-EM map of RNF168/UbcH5c-Ub/nucleosome determined by E2-Ub-NCP conjugation strategy
Method: single particle / : Ai H, Zebin T, Deng Z, Pan M, Liu L

EMDB-44482: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab
Method: single particle / : Gorman J, Kwong PD
EMDB-44484: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs
Method: single particle / : Gorman J, Kwong PD
EMDB-44491: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab
Method: single particle / : Gorman J, Kwong PD

PDB-9ber: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab
Method: single particle / : Gorman J, Kwong PD

PDB-9bew: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs
Method: single particle / : Gorman J, Kwong PD

PDB-9bf6: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab
Method: single particle / : Gorman J, Kwong PD

EMDB-38873: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38874: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38875: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38876: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y36: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y37: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y38: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y39: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-43046: 
Full length integrin AlphaIIbBeta3 in pre-active state
Method: single particle / : Huo T, Wu H, Wang Z

EMDB-32979: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)
Method: single particle / : Zheng Q, Zhu R, Sun H, Cheng T, Li S, Xia N

PDB-7x35: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)
Method: single particle / : Zheng Q, Zhu R, Sun H, Cheng T, Li S, Xia N

EMDB-50409: 
Subtomogram average of 80S ribosomes in native S. cerevisiae
Method: subtomogram averaging / : Spindler MC, Mahamid J

EMDB-50415: 
Subtomogram average of 80S ribosomes in S. cerevisiae under acute glucose starvation
Method: subtomogram averaging / : Spindler MC, Mahamid J

EMDB-38617: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38618: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38619: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38620: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38621: 
SARS-CoV-2 spike + IMCAS-123
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38823: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex
Method: single particle / : Jia GW, Tong Z, Tong JY, Su ZM

PDB-8xse: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsf: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsi: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsj: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsl: 
SARS-CoV-2 spike + IMCAS-123
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8y0y: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex
Method: single particle / : Jia GW, Tong Z, Tong JY, Su ZM

EMDB-38860: 
structure of RSF-147bp NCP complex Class 0
Method: single particle / : Zhang JL

EMDB-38861: 
Structure of RSF-147bpNCP complex class 2
Method: single particle / : Zhang JL

EMDB-38865: 
RSF-38N38NCP complex Class 2
Method: single particle / : Zhang JL

EMDB-37895: 
PNPase mutant of Mycobacterium tuberculosis
Method: single particle / : Wang N, Sheng YN, Liu YT

EMDB-37896: 
PNPase of M.tuberculosis with its RNA substrate
Method: single particle / : Wang N, Sheng YN

EMDB-37905: 
PNPase of Mycobacterium tuberculosis
Method: single particle / : Wang N, Sheng YN, Liu YT

PDB-8wwp: 
PNPase mutant of Mycobacterium tuberculosis
Method: single particle / : Wang N, Sheng YN, Liu YT

PDB-8wx0: 
PNPase of M.tuberculosis with its RNA substrate
Method: single particle / : Wang N, Sheng YN
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
