-Search query
-Search result
Showing 1 - 50 of 3,124 items for (author: tian & x)

EMDB-31367: 
Structure of mumps virus nucleoprotein without C-arm
Method: helical / : Shen Q, Shan H, Zhang N, Qin Y
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
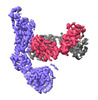
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
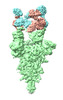
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
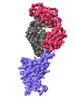
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
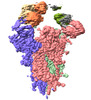
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-38694: 
Human TOM complex with whole Tom20
Method: single particle / : Tian XY, Su JY, Sui SF

EMDB-60277: 
TOM core complex
Method: single particle / : Tian XY, Su JY, Sui SF

EMDB-60278: 
whole Tom20
Method: single particle / : Tian XY, Su JY, Sui SF
EMDB-38099: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by intein-based E2-Ub-NCP conjugation strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
EMDB-38100: 
Cryo-EM structures of RNF168/UbcH5c-Ub/nucleosomes complex determined by activity-based chemical trapping strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
EMDB-38101: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by activity-based chemical trapping strategy (adjacent H2AK13/15 dual-monoubiquitination)
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
EMDB-38102: 
Cryo-EM map of RNF168/UbcH5c-Ub/nucleosome determined by E2-Ub-NCP conjugation strategy
Method: single particle / : Ai H, Zebin T, Deng Z, Pan M, Liu L

EMDB-36914: 
Cryo-EM structure of Streptomyces coelicolor transcription initiation complex with the global transcription factor AfsR
Method: single particle / : Lin W, Shi J, Xu JC

EMDB-37515: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.
Method: single particle / : Tan J, Xiao Y, Kong F, Lei J, Yuan Y, Yan C

EMDB-37520: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.
Method: single particle / : Tan J, Xiao Y, Kong F, Lei J, Yuan Y, Yan C

EMDB-39729: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.
Method: single particle / : Tan J, Xiao Y, Kong F, Lei J, Yuan Y, Yan C

PDB-8wgr: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.
Method: single particle / : Tan J, Xiao Y, Kong F, Lei J, Yuan Y, Yan C

PDB-8wgx: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.
Method: single particle / : Tan J, Xiao Y, Kong F, Lei J, Yuan Y, Yan C

PDB-8z1l: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.
Method: single particle / : Tan J, Xiao Y, Kong F, Lei J, Yuan Y, Yan C

EMDB-38873: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38874: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38875: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38876: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y36: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y37: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y38: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y39: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-43762: 
Aca2 from Pectobacterium phage ZF40 bound to RNA
Method: single particle / : Wilkinson ME, Birkholz N, Kimanius D, Fineran PC

PDB-8w35: 
Aca2 from Pectobacterium phage ZF40 bound to RNA
Method: single particle / : Wilkinson ME, Birkholz N, Kimanius D, Fineran PC

EMDB-60607: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-60608: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Ggust
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-60626: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH
EMDB-60627: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
